Calibration-Aimed Comparison of Image-Cytometry- and Flow-Cytometry-Based Approaches of Ploidy Analysis
- PMID: 36146303
- PMCID: PMC9502733
- DOI: 10.3390/s22186952
Calibration-Aimed Comparison of Image-Cytometry- and Flow-Cytometry-Based Approaches of Ploidy Analysis
Abstract
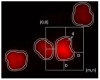
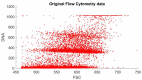
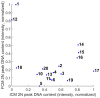
Ploidy analysis is the fundamental method of measuring DNA content. For decades, the principal way of conducting ploidy analysis was through flow cytometry. A flow cytometer is a specialized tool for analyzing cells in a solution. This is convenient in laboratory environments, but prohibits measurement reproducibility and the complete detachment of sample preparation from data acquisition and analysis, which seems to have become paramount with the constant decrease in the number of pathologists per capita all over the globe. As more open computer-aided systems emerge in medicine, the demand for overcoming these shortcomings, and opening access to even more (and more flexible) options, has also emerged. Image-based analysis systems can provide an alternative to these types of workloads, placing the abovementioned problems in a different light. Flow cytometry data can be used as a reference for calibrating an image-based system. This article aims to show an approach to constructing an image-based solution for ploidy analysis, take measurements for a basic comparison of the data produced by the two methods, and produce a workflow with the ultimate goal of calibrating the image-based system.
Keywords: DNA content; calibration; cytology; cytometry; digital pathology; ploidy analysis.
Conflict of interest statement
The authors declare no conflict of interest.
Figures












References
-
- Krenács T., Zsakovics I., Micsik T., Fonyad L., Varga V., Ficsor L., Kiszler G., Molnár B. Digital Microscopy: The Upcoming Revolution in Histopathology Teaching, Diagnostics, Research and Quality Assurance 2010. [(accessed on 7 August 2020)]. Available online: https://www.researchgate.net/publication/232607690.
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Research Materials

