Next-generation sequencing based detection of BRCA1 and BRCA2 large genomic rearrangements in Chinese cancer patients
- PMID: 36147908
- PMCID: PMC9487528
- DOI: 10.3389/fonc.2022.898916
Next-generation sequencing based detection of BRCA1 and BRCA2 large genomic rearrangements in Chinese cancer patients
Abstract
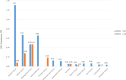
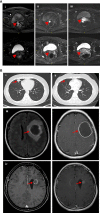
BRCA1/2 mutation is a biomarker for guiding multiple solid tumor treatment. However, the prevalence of BRCA1/2 large genomic rearrangements (LGRs) in Chinese cancer patients has not been well revealed partially due to technical difficulties in LGR detection. This study utilized next-generation sequencing (NGS) to analyze the BRCA1/2 mutation profile, including LGR, in 56126 Chinese cancer patients. We also reported that two ovarian and breast cancer patients with NGS-determined BRCA1/2 LGR benefited from PARP inhibitors (PARPi). DNA sequencing identified BRCA1/2 variants (including LGR, pathogenic and likely-pathogenic variants) in 2108 individuals. Seventy patients were discovered to harbor germline LGRs in BRCA1 and 14 had germline LGRs in BRCA2. Among the LGRs detected, exon 1-2 deletion was the predominant LGR (14/70) in BRCA1, and exon 22-24 deletion was the most frequent LGR (3/14) in BRCA2. Notably, the BRCA1 exon 7 deletion was a novel LGR and was identified in six patients, suggesting a specific LGR in Chinese cancer patients. The prevalence analysis of BRCA1 and BRCA2 LGRs across multiple cancers revealed that BRCA1 LGR more frequently occurred in ovarian cancer (1.31%, 33/2526), and BRCA2 LGR was more commonly seen in cholangiocarcinoma (0.47%, 2/425). Two ovarian and breast cancer patients with BRCA1/2 LGR benefited from PARPi therapy. This is the first study to reveal the BRCA1/2 LGR profile of a Chinese pan-cancer cohort by using an NGS-based assay. Two breast and ovarian cancer patients harboring NGS-determined BRCA1/2 LGR benefited from PARPi, indicating that NGS-based detection of BRCA1/2 LGR has the potential to guide PARPi treatment.
Keywords: BRCA; breast cancer; large genomic rearrangements; next-generation sequencing; ovarian cancer; pan-cancer.
Copyright © 2022 Hua, Tian, Wang, Bei, Cui, Zhang, Bao, Bai, Zhao and Yuan.
Conflict of interest statement
D Hua, T Bei, L Cui, B Zhang, C Bao, Y Bai, and X Zhao are employees of 3D Medicines Inc. The remaining authors declare that the research was conducted in the absence of any commercial or financial relationships that could be constructed as a potential conflict of interest. The reviewer (JL) declared a shared affiliation, with no collaboration, with the authors (XW, PY) to the handling editor at the time of review
Figures

Similar articles
-
Complex Characterization of Germline Large Genomic Rearrangements of the BRCA1 and BRCA2 Genes in High-Risk Breast Cancer Patients-Novel Variants from a Large National Center.Int J Mol Sci. 2020 Jun 30;21(13):4650. doi: 10.3390/ijms21134650. Int J Mol Sci. 2020. PMID: 32629901 Free PMC article.
-
Comprehensive mutation detection of BRCA1/2 genes reveals large genomic rearrangements contribute to hereditary breast and ovarian cancer in Chinese women.BMC Cancer. 2019 Jun 7;19(1):551. doi: 10.1186/s12885-019-5765-3. BMC Cancer. 2019. PMID: 31174498 Free PMC article.
-
Large genomic rearrangements in the familial breast and ovarian cancer gene BRCA1 are associated with an increased frequency of high risk features.Fam Cancer. 2015 Jun;14(2):287-95. doi: 10.1007/s10689-015-9785-0. Fam Cancer. 2015. PMID: 25678442
-
[Detecting Large Germline Rearrangements of BRCA1 by Next Generation Tumor Sequencing].Mol Biol (Mosk). 2020 May-Jun;54(4):688-698. doi: 10.31857/S0026898420040114. Mol Biol (Mosk). 2020. PMID: 32840490 Review. Russian.
-
Detection of BRCA1/2 large genomic rearrangements in breast and ovarian cancer patients: an overview of the current methods.Expert Rev Mol Diagn. 2019 Sep;19(9):795-802. doi: 10.1080/14737159.2019.1657011. Epub 2019 Aug 28. Expert Rev Mol Diagn. 2019. PMID: 31429350 Review.
Cited by
-
Population-based BRCA germline mutation screening in the Han Chinese identifies individuals at risk of BRCA mutation-related cancer: experience from a clinical diagnostic center from greater Shanghai area.BMC Cancer. 2024 Apr 2;24(1):411. doi: 10.1186/s12885-024-12089-w. BMC Cancer. 2024. PMID: 38566028 Free PMC article.
-
Comprehensive profiling of pathogenic germline large genomic rearrangements in a pan-cancer analysis.Mol Oncol. 2023 Sep;17(9):1917-1929. doi: 10.1002/1878-0261.13430. Epub 2023 Apr 12. Mol Oncol. 2023. PMID: 37013911 Free PMC article.
-
Mutational spectrum of BRCA genes in Egyptian patients with breast cancer.Sci Rep. 2025 Jul 18;15(1):26067. doi: 10.1038/s41598-025-09810-5. Sci Rep. 2025. PMID: 40681599 Free PMC article.
References
-
- (NCCN) . N.C.C.N., NCCN clinical practice guidelines in oncology. In: Ovarian cancer. version 1. 2021. 3025 Chemical Road, Suite 100, Plymouth Meeting, PA 19462, USA: National Comprehensive Cancer Network (NCCN; (2021).
-
- (NCCN) . N.C.C.N., NCCN clinical practice guidelines in oncology. In: Breast cancer. version 4. 2021. 3025 Chemical Road, Suite 100, Plymouth Meeting, PA 19462, USA: National Comprehensive Cancer Network (NCCN; (2021).
-
- Bozsik A, Pocza T, Papp J, Vaszko T, Butz H, Patocs A, et al. . Complex characterization of germline Large genomic rearrangements of the BRCA1 and BRCA2 genes in high-risk breast cancer patients-novel variants from a Large national center. Int J Mol Sci (2020) 21(13):4605. doi: 10.3390/ijms21134650 - DOI - PMC - PubMed
-
- Germani A, Libi F, Maggi S, Stanzani G, Lombardi A, Pellegrini P, et al. . Rapid detection of copy number variations and point mutations in BRCA1/2 genes using a single workflow by ion semiconductor sequencing pipeline. Oncotarget (2018) 9(72):33648–55. doi: 10.18632/oncotarget.26000 - DOI - PMC - PubMed
LinkOut - more resources
Full Text Sources
Miscellaneous

