Tumor Cell Extrinsic Synaptogyrin 3 Expression as a Diagnostic and Prognostic Biomarker in Head and Neck Cancer
- PMID: 36148399
- PMCID: PMC9491693
- DOI: 10.1158/2767-9764.crc-21-0135
Tumor Cell Extrinsic Synaptogyrin 3 Expression as a Diagnostic and Prognostic Biomarker in Head and Neck Cancer
Abstract
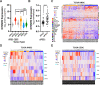
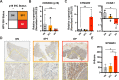
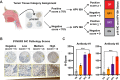
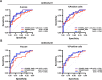
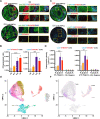
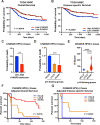
Over 70% of oropharyngeal head and neck squamous cell carcinoma (HNSC) cases in the United States are positive for human papillomavirus (HPV) yet biomarkers for stratifying oropharyngeal head and neck squamous cell carcinoma (HNSC) patient risk are limited. We used immunogenomics to identify differentially expressed genes in immune cells of HPV(+) and HPV(-) squamous carcinomas. Candidate genes were tested in clinical specimens using both quantitative RT-PCR and IHC and validated by IHC using the Carolina Head and Neck Cancer Study (CHANCE) tissue microarray of HNSC cases. We performed multiplex immunofluorescent staining to confirm expression within the immune cells of HPV(+) tumors, receiver operating characteristic (ROC) curve analyses, and assessed survival outcomes. The neuronal gene Synaptogyrin-3 (SYNGR3) is robustly expressed in immune cells of HPV(+) squamous cancers. Multiplex immunostaining and single cell RNA-seq analyses confirmed SYNGR3 expression in T cells, but also unexpectedly in B cells of HPV(+) tumors. ROC curve analyses revealed that combining SYNGR3 and p16 provides more sensitivity and specificity for HPV detection compared to p16 IHC alone. SYNGR3-high HNSC patients have significantly better prognosis with five-year OS and DSS rates of 60% and 71%, respectively. Moreover, combining p16 localization and SYNGR3 expression can further risk stratify HPV(+) patients such that high cytoplasmic, low nuclear p16 do significantly worse (Hazard Ratio, 8.6; P = 0.032) compared to patients with high cytoplasmic, high nuclear p16. SYNGR3 expression in T and B cells is associated with HPV status and enhanced survival outcomes of HNSC patients.
Keywords: Diagnostic biomarkers; HPV(+) HNSC; Head; Neck & Oral Cancers; Prognostic biomarkers; Tumor Immune Microenvironment biomarker.
Conflict of interest statement
R.M. Murphy reports grants from NIH NIDCR during the conduct of the study. S. Kumar reports a patent to Method for quantifying DNA fragments in a sample by size issued, licensed, and with royalties paid and a patent to Compositions and methods for the selective detection of tumor-derived viral DNA issued, licensed, and with royalties paid. P.Y.F. Zeng reports a patent to prognosticate HPV+ head and neck cancer patients pending. D.N. Hayes reports grants from NCI during the conduct of the study. G.P. Gupta reports personal fees and other from Naveris outside the submitted work; in addition, G.P. Gupta has a patent to detect tumor-modified HPV in blood issued, licensed, and with royalties paid. C.V. Pecot reports a patent to US Provisional 63/215,674 pending. A.L. Amelio reports personal fees from LG Life Sciences outside the submitted work; in addition, A.L. Amelio reports a patent to USPTO Provisional 63/215,674 pending. No other disclosures were reported.
Figures
Similar articles
-
Quantitative assessment of p16 expression in FNA specimens from head and neck squamous cell carcinoma and correlation with HPV status.Cancer Cytopathol. 2021 May;129(5):394-404. doi: 10.1002/cncy.22399. Epub 2020 Dec 28. Cancer Cytopathol. 2021. PMID: 33369885 Free PMC article.
-
Heterogeneity of p16 immunohistochemistry and increased sensitivity of RNA in situ hybridization in cytology specimens of HPV-related head and neck squamous cell carcinoma.Cancer Cytopathol. 2019 Oct;127(10):632-642. doi: 10.1002/cncy.22178. Epub 2019 Sep 11. Cancer Cytopathol. 2019. PMID: 31509355
-
A novel prediction model for human papillomavirus-associated oropharyngeal squamous cell carcinoma using p16 and subcellular β-catenin expression.J Oral Pathol Med. 2016 Jul;45(6):399-408. doi: 10.1111/jop.12378. Epub 2015 Oct 22. J Oral Pathol Med. 2016. PMID: 26493274 Free PMC article.
-
The Next Chapter in Cancer Diagnostics: Advances in HPV-Positive Head and Neck Cancer.Biomolecules. 2024 Jul 30;14(8):925. doi: 10.3390/biom14080925. Biomolecules. 2024. PMID: 39199313 Free PMC article. Review.
-
HPV-related squamous cell carcinoma of the head and neck: An update on testing in routine pathology practice.Semin Diagn Pathol. 2015 Sep;32(5):344-51. doi: 10.1053/j.semdp.2015.02.013. Epub 2015 Feb 4. Semin Diagn Pathol. 2015. PMID: 25724476 Review.
Cited by
-
The value of lymphocyte to monocyte ratio in the prognosis of head and neck squamous cell carcinoma: a meta-analysis.PeerJ. 2023 Sep 11;11:e16014. doi: 10.7717/peerj.16014. eCollection 2023. PeerJ. 2023. PMID: 37719125 Free PMC article.
References
-
- Bray F, Ferlay J, Soerjomataram I, Siegel RL, Torre LA, Jemal A. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin 2018;68:394–424. - PubMed
-
- Global Burden of Disease Cancer Collaboration; Fitzmaurice C, Allen C, Barber RM, Barregard L, Bhutta ZA, et al. . Global, regional, and national cancer incidence, mortality, years of life lost, years lived with disability, and disability-adjusted life-years for 32 cancer groups, 1990 to 2015: a systematic analysis for the global burden of disease study. JAMA Oncol 2016;3:524–48. - PMC - PubMed
-
- Siegel RL, Miller KD, Fuchs HE, Jemal A. Cancer statistics, 2021. CA Cancer J Clin 2021;71:7–33. - PubMed
-
- Sacco AG, Cohen EE. Current treatment options for recurrent or metastatic head and neck squamous cell carcinoma. J Clin Oncol 2015;33:3305–15. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases

