Novel pyroptosis-associated genes signature for predicting the prognosis of sarcoma and validation
- PMID: 36155774
- PMCID: PMC9742515
- DOI: 10.1042/BSR20221053
Novel pyroptosis-associated genes signature for predicting the prognosis of sarcoma and validation
Abstract
Background: Sarcoma is a rare mesenchymal malignant tumor. Recently, pyroptosis has been reported to be a mode of programmed cell death. Nonetheless, levels of pyroptosis-associated genes in sarcoma and its relevance to prognostic outcomes are yet to be elucidated.
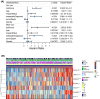
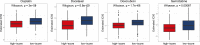
Results: Sarcoma cases were classified into two subtypes with regards to differentially expressed genes. We established a profile composed of seven genes and classified the sarcoma patients into low- and high-risk groups through least absolute shrinkage and selection operator Cox regression. Survival rate of low-risk sarcoma patients was markedly higher, relative to high-risk group (P<0.001). In combination with clinical features, the risk score was established to be an independent predictive factor for OS of sarcoma patients. Chemotherapeutic drug sensitivity response analysis found 65 drugs with higher drug sensitivity in low-risk, than in high-risk group and 14 drugs with higher drug sensitivity in the high-risk patient group, compared with low-risk patient group. In addition, functional enrichment, pathway and gene mutation of the two modules were analyzed. Finally, we used qRT-PCR to detect the expression of seven pyroptosis-related genes in tumor cells, and human skeletal muscle cells, compared with human skeletal muscle cells, PODXL2, LRRC17, GABRA3, SCUBE3 and RFLNB genes show high expression levels in tumor cells, while IGHG2 and hepatic leukemia factor show low expression levels in tumor cells.
Conclusions: Our research suggest that pyroptosis is closely associated with sarcoma, and these findings confirm that pyroptosis-associated seven genes have a critical role in sarcoma and are potential prognostic factors for sarcoma.
Keywords: biomarkers; pyroptosis; sacoma.
© 2022 The Author(s).
Conflict of interest statement
The authors declare that there are no competing interests associated with the manuscript.
Figures





Similar articles
-
Construction and Validation of a Novel Pyroptosis-Related Four-lncRNA Prognostic Signature Related to Gastric Cancer and Immune Infiltration.Front Immunol. 2022 Mar 22;13:854785. doi: 10.3389/fimmu.2022.854785. eCollection 2022. Front Immunol. 2022. PMID: 35392086 Free PMC article.
-
Pyroptosis, apoptosis, and necroptosis molecular subtype derived prognostic signature universal applicable for gastric cancer-A large sample and multicenter retrospective analysis.Comput Biol Med. 2022 Oct;149:106037. doi: 10.1016/j.compbiomed.2022.106037. Epub 2022 Aug 27. Comput Biol Med. 2022. PMID: 36044785
-
Pyroptosis-Related Gene Signature Is a Novel Prognostic Biomarker for Sarcoma Patients.Dis Markers. 2021 Dec 3;2021:9919842. doi: 10.1155/2021/9919842. eCollection 2021. Dis Markers. 2021. PMID: 34904022 Free PMC article.
-
Identification of a pyroptosis-related prognostic signature in breast cancer.BMC Cancer. 2022 Apr 20;22(1):429. doi: 10.1186/s12885-022-09526-z. BMC Cancer. 2022. PMID: 35443644 Free PMC article.
-
A Novel Risk Signature with Seven Pyroptosis-Related Genes for Prognosis Prediction in Glioma.World Neurosurg. 2022 Mar;159:e285-e302. doi: 10.1016/j.wneu.2021.12.042. Epub 2021 Dec 18. World Neurosurg. 2022. PMID: 34929369
Cited by
-
Serum Proteomic Profiling in Patients with Chronic Obstructive Pulmonary Disease.Int J Chron Obstruct Pulmon Dis. 2023 Jul 28;18:1623-1635. doi: 10.2147/COPD.S413924. eCollection 2023. Int J Chron Obstruct Pulmon Dis. 2023. PMID: 37533772 Free PMC article.
-
Microwave ablation of non-small cell lung cancer enhances local T-cell abundance and alters monocyte interactions.BMC Cancer. 2025 Apr 3;25(1):605. doi: 10.1186/s12885-025-14002-5. BMC Cancer. 2025. PMID: 40181307 Free PMC article.
References
-
- JGeorge D Demetri 1, Scott Antonia, Robert S Benjamin, Marilyn M Bui, Ephraim S Casper, et al. (2010) Soft tissue sarcoma. J Natl Compr Canc Netw 8, 6, 630–674 - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Miscellaneous

