Single cell atlas of spinal cord injury in mice reveals a pro-regenerative signature in spinocerebellar neurons
- PMID: 36163250
- PMCID: PMC9513082
- DOI: 10.1038/s41467-022-33184-1
Single cell atlas of spinal cord injury in mice reveals a pro-regenerative signature in spinocerebellar neurons
Abstract
After spinal cord injury, tissue distal to the lesion contains undamaged cells that could support or augment recovery. Targeting these cells requires a clearer understanding of their injury responses and capacity for repair. Here, we use single nucleus RNA sequencing to profile how each cell type in the lumbar spinal cord changes after a thoracic injury in mice. We present an atlas of these dynamic responses across dozens of cell types in the acute, subacute, and chronically injured spinal cord. Using this resource, we find rare spinal neurons that express a signature of regeneration in response to injury, including a major population that represent spinocerebellar projection neurons. We characterize these cells anatomically and observed axonal sparing, outgrowth, and remodeling in the spinal cord and cerebellum. Together, this work provides a key resource for studying cellular responses to injury and uncovers the spontaneous plasticity of spinocerebellar neurons, uncovering a potential candidate for targeted therapy.
© 2022. This is a U.S. Government work and not under copyright protection in the US; foreign copyright protection may apply.
Conflict of interest statement
The authors declare the following competing financial interests: G.C. is a consultant with ONWARD Medical. G.C. is a shareholder of ONWARD Medical, a company developing an EES-based therapy to restore movement after spinal cord injury. The remaining authors declare no competing interests.
Figures




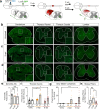
References
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases
Research Materials

