BMPR1A promotes ID2-ZEB1 interaction to suppress excessive endothelial to mesenchymal transition
- PMID: 36166408
- PMCID: PMC10409893
- DOI: 10.1093/cvr/cvac159
BMPR1A promotes ID2-ZEB1 interaction to suppress excessive endothelial to mesenchymal transition
Abstract
Aims: Components of bone morphogenetic protein (BMP) signalling have been implicated in both pathogenesis of pulmonary arterial hypertension (PAH) and endothelial-mesenchymal transition (EndoMT). In particular, the importance of BMP type 2 receptor in these processes has been extensively analysed. However, the contribution of BMP type 1 receptors (BMPR1s) to the onset of PAH and EndoMT remains poorly understood. BMPR1A, one of BMPR1s, was recently implicated in the pathogenesis of PAH, and was found to be down-regulated in the lungs of PAH patients, neither the downstream mechanism nor its contribution to EndoMT has been described. Therefore, we aim to delineate the role of endothelial BMPR1A in modulating EndoMT and pathogenesis of PAH.
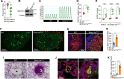
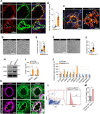
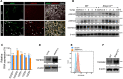
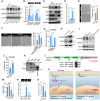
Methods and results: We find that BMPR1A knockdown in endothelial cells (ECs) induces hallmarks of EndoMT, and deletion of endothelial Bmpr1a in adult mice (Bmpr1aiECKO) leads to development of PAH-like symptoms due to excessive EndoMT. By lineage tracing, we show that endothelial-derived smooth muscle cells are increased in endothelial Bmpr1a-deleted mice. Mechanistically, we identify ZEB1 as a primary target for BMPR1A in this setting; upon BMPR1A activation, ID2 physically interacts and sequesters ZEB1 to attenuate transcription of Tgfbr2, which in turn lowers the responses of ECs towards transforming growth factor beta (TGFβ) stimulation and prevents excessive EndoMT. In Bmpr1aiECKO mice, administering endothelial targeting lipid nanoparticles containing siRNA against Tgfbr2 effectively ameliorate PAH, reiterating the importance of BMPR1A-ID2/ZEB1-TGFBR2 axis in modulating progression of EndoMT and pathogenesis of PAH.
Conclusions: We demonstrate that BMPR1A is key to maintain endothelial identity and to prevent excessive EndoMT. We identify BMPR1A-induced interaction between ID2 and ZEB1 is the key regulatory step for onset of EndoMT and pathogenesis of PAH. Our findings indicate that BMPR1A-ID2/ZEB1-TGFBR2 signalling axis could serve as a potential novel therapeutic target for PAH and other EndoMT-related vascular disorders.
Keywords: BMP signalling; BMPR1A; EndoMT; Pulmonary arterial hypertension; Vascular remodelling.
© The Author(s) 2022. Published by Oxford University Press on behalf of the European Society of Cardiology. All rights reserved. For permissions, please email: journals.permissions@oup.com.
Conflict of interest statement
Conflict of interest: None declared.
Figures

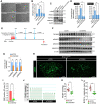
References
-
- Farber HW, Loscalzo J. Pulmonary arterial hypertension. N Engl J Med 2004;351:1655–1665. - PubMed
-
- Michelakis ED, Wilkins MR, Rabinovitch M. Emerging concepts and translational priorities in pulmonary arterial hypertension. Circulation 2008;118:1486–1495. - PubMed
-
- Evans JD, Girerd B, Montani D, Wang XJ, Galie N, Austin ED, Elliott G, Asano K, Grunig E, Yan Y, Jing ZC, Manes A, Palazzini M, Wheeler LA, Nakayama I, Satoh T, Eichstaedt C, Hinderhofer K, Wolf M, Rosenzweig EB, Chung WK, Soubrier F, Simonneau G, Sitbon O, Graf S, Kaptoge S, Di Angelantonio E, Humbert M, Morrell NW. BMPR2 Mutations and survival in pulmonary arterial hypertension: an individual participant data meta-analysis. Lancet Respir Med 2016;4:129–137. - PMC - PubMed
-
- Du L, Sullivan CC, Chu D, Cho AJ, Kido M, Wolf PL, Yuan JX, Deutsch R, Jamieson SW, Thistlethwaite PA. Signaling molecules in nonfamilial pulmonary hypertension. N Engl J Med 2003;348:500–509. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Research Materials

