BMSC-derived exosomal miR-27a-3p and miR-196b-5p regulate bone remodeling in ovariectomized rats
- PMID: 36168439
- PMCID: PMC9509671
- DOI: 10.7717/peerj.13744
BMSC-derived exosomal miR-27a-3p and miR-196b-5p regulate bone remodeling in ovariectomized rats
Abstract
Background: In the bone marrow microenvironment of postmenopausal osteoporosis (PMOP), bone marrow mesenchymal stem cell (BMSC)-derived exosomal miRNAs play an important role in bone formation and bone resorption, although the pathogenesis has yet to be clarified.
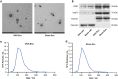
Methods: BMSC-derived exosomes from ovariectomized rats (OVX-Exo) and sham-operated rats (Sham-Exo) were co-cultured with bone marrow-derived macrophages to study their effects on osteoclast differentiation. Next-generation sequencing was utilized to identify the differentially expressed miRNAs (DE-miRNAs) between OVX-Exo and Sham-Exo, while target genes were analyzed using bioinformatics. The regulatory effects of miR-27a-3p and miR-196b-5p on osteogenic differentiation of BMSCs and osteoclast differentiation were verified by gain-of-function and loss-of-function analyses.
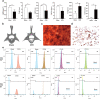
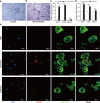
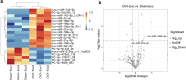
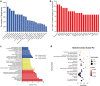
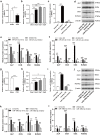
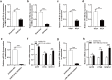
Results: Osteoclast differentiation was significantly enhanced in the OVX-Exo treatment group compared to the Sham-Exo group. Twenty DE-miRNAs were identified between OVX-Exo and Sham-Exo, among which miR-27a-3p and miR-196b-5p promoted the expressions of osteogenic differentiation markers in BMSCs. In contrast, knockdown of miR-27a-3p and miR-196b-5p increased the expressions of osteoclastic markers in osteoclast. These 20 DE-miRNAs were found to target 11435 mRNAs. Gene Ontology and Kyoto Encyclopedia of Genes and Genomes pathway analyses revealed that these target genes were involved in several biological processes and osteoporosis-related signaling pathways.
Conclusion: BMSC-derived exosomal miR-27a-3p and miR-196b-5p may play a positive regulatory role in bone remodeling.
Keywords: Bone marrow mesenchymal stem cell; Exosomes; MicroRNAs; Next-generation sequencing; Osteoclast; Osteogenic differentiation; Postmenopausal osteoporosis.
©2022 Lai et al.
Conflict of interest statement
The authors declare there are no competing interests.
Figures
Similar articles
-
Tumour necrosis factor α regulates the miR-27a-3p-Sfrp1 axis in a mouse model of osteoporosis.Exp Physiol. 2024 Jul;109(7):1109-1123. doi: 10.1113/EP090311. Epub 2024 May 15. Exp Physiol. 2024. PMID: 38748896 Free PMC article.
-
Plasma exosomal miR-30a-5p inhibits osteogenic differentiation of bone marrow mesenchymal stem cells from a chronic unpredictable mild stress-induced depression rat model.Mol Cell Probes. 2024 Jun;75:101957. doi: 10.1016/j.mcp.2024.101957. Epub 2024 Mar 23. Mol Cell Probes. 2024. PMID: 38513992
-
LncRNA SNHG14 Delivered by Bone Marrow Mesenchymal Stem Cells-Secreted Exosomes Regulates Osteogenesis and Adipogenesis in Osteoporosis by Mediating the miR-27a-3p/LMNB1 Axis.Kaohsiung J Med Sci. 2025 May;41(5):e70004. doi: 10.1002/kjm2.70004. Epub 2025 Mar 7. Kaohsiung J Med Sci. 2025. PMID: 40052307 Free PMC article.
-
Exosomal Osteoclast-Derived miRNA in Rheumatoid Arthritis: From Their Pathogenesis in Bone Erosion to New Therapeutic Approaches.Int J Mol Sci. 2024 Jan 25;25(3):1506. doi: 10.3390/ijms25031506. Int J Mol Sci. 2024. PMID: 38338785 Free PMC article. Review.
-
[Exercise regulates bone metabolism via microRNAs].Sheng Li Xue Bao. 2023 Jun 25;75(3):429-438. Sheng Li Xue Bao. 2023. PMID: 37340651 Review. Chinese.
Cited by
-
Advances in the Study of Exosomes as Drug Delivery Systems for Bone-Related Diseases.Pharmaceutics. 2023 Jan 9;15(1):220. doi: 10.3390/pharmaceutics15010220. Pharmaceutics. 2023. PMID: 36678850 Free PMC article. Review.
-
Identification and evaluation of circulating exosomal miRNAs for the diagnosis of postmenopausal osteoporosis.J Orthop Surg Res. 2023 Jul 26;18(1):533. doi: 10.1186/s13018-023-04020-z. J Orthop Surg Res. 2023. PMID: 37496029 Free PMC article.
-
Macrophages-bone marrow mesenchymal stem cells crosstalk in bone healing.Front Cell Dev Biol. 2023 Jun 23;11:1193765. doi: 10.3389/fcell.2023.1193765. eCollection 2023. Front Cell Dev Biol. 2023. PMID: 37427382 Free PMC article. Review.
-
Human umbilical cord mesenchymal stem cells derived extracellular vesicles alleviate salpingitis by promoting M1-to-M2 transformation.Front Physiol. 2023 Feb 16;14:1131701. doi: 10.3389/fphys.2023.1131701. eCollection 2023. Front Physiol. 2023. PMID: 36875046 Free PMC article.
-
Correlation Between miR-27a-3p Polymorphisms and Peri-Implantitis Susceptibility: A Case-Control Study.Int Dent J. 2025 Jun;75(3):1921-1928. doi: 10.1016/j.identj.2025.01.013. Epub 2025 Feb 10. Int Dent J. 2025. PMID: 39934011 Free PMC article.
References
-
- Furesi G, De Jesus Domingues AM, Alexopoulou D, Dahl A, Hackl M, Schmidt JR, Kalkhof S, Kurth T, Taipaleenmäki H, Conrad S, Hofbauer C, Rauner M, Hofbauer LC. Exosomal miRNAs from prostate cancer impair osteoblast function in mice. International Journal of Molecular Sciences. 2022;23:1285. doi: 10.3390/ijms23031285. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

