FOXP3+ macrophage represses acute ischemic stroke-induced neural inflammation
- PMID: 36170234
- PMCID: PMC10012925
- DOI: 10.1080/15548627.2022.2116833
FOXP3+ macrophage represses acute ischemic stroke-induced neural inflammation
Abstract
Proper termination of cell-death-induced neural inflammation is the premise of tissue repair in acute ischemic stroke (AIS). Macrophages scavenge cell corpses/debris and produce inflammatory mediators that orchestrate immune responses. Here, we report that FOXP3, the key immune-repressive transcription factor of Tregs, is conditionally expressed in macrophages in stroke lesion. FOXP3 ablation in macrophages results in detrimental stroke outcomes, emphasizing the beneficial role of FOXP3+ macrophages. FOXP3+ macrophages are distinct from the M1 or M2 subsets and display superactive efferocytic capacity. With scRNAseq and analysis of FOXP3-bound-DNA isolated with CUT & RUN, we show that FOXP3 facilitates macrophage phagocytosis through enhancing cargo metabolism. FOXP3 expression is controlled by macroautophagic/autophagic protein degradation in resting macrophages, while initiation of LC3-associated phagocytosis (LAP) competitively occupies the autophagic machineries, and thus permits FOXP3 activation. Our data demonstrate a distinct set of FOXP3+ macrophages with enhanced scavenging capability, which could be a target in immunomodulatory therapy against AIS.Abbreviations: ADGRE1/F4/80: adhesion G protein-coupled receptor E1; AIF1/Iba1: allograft inflammatory factor 1; AIS: acute ischemic stroke; ARG1: arginase 1; ATP: adenosine triphosphate; BECN1/Beclin1: Beclin 1, autophagy related; BMDM: bone marrow-derived macrophages; CKO: conditional knockout; CSF1/M-CSF: colony stimulating factor 1 (macrophage); CSF2/GM-CSF: colony stimulating factor 2; CSF3/G-CSF: colony stimulating factor 3; CUT & RUN: cleavage under targets and release using nuclease; CyD: cytochalasin D; DAMP: danger/damage-associated molecular pattern; DIL: dioctadecyl-3,3,3,3-tetramethylin docarbocyanine; ELISA: enzyme linked immunosorbent assay; GO: Gene Ontology; FCGR3/CD16: Fc receptor, IgG, low affinity III; HMGB1: high mobility group box 1; IFNG/IFNγ: interferon gamma; IP: immunoprecipitation; KEGG: Kyoto Encyclopedia of Genes and Genomes; ITGAM/CD11b: integrin subunit alpha M; ITGAX/CD11c: integrin subunit alpha X; LAP: LC3-associated phagocytosis; LC-MS: liquid chromatography-mass spectrometry; LPS: lipopolysaccharide; MRC1/CD206: mannose receptor, C type 1; O4: oligodendrocyte marker O4; PBMC: peripheral blood mononuclear cells; RBC: red blood cells; PTPRC/CD45: protein tyrosine phosphatase, receptor type, C; RBFOX3/NeuN: RNA binding protein, fox 1 homolog (C. elegans) 3; RUBCN/Rubicon: RUN domain and cysteine-rich domain containing, Beclin 1-interacting protein; scRNAseq: single cell RNA sequencing; SQSTM1/p62 (sequestosome 1); TGFB/TGFβ: transforming growth factor, beta; tMCAO: transient middle cerebral artery occlusion; TNF/TNFα: tumor necrosis factor; Treg: regulatory T cell.
Keywords: FOXP3; inflammation; macrophage; phagocytosis; stroke.
Conflict of interest statement
No potential conflict of interest was reported by the author(s).
Figures
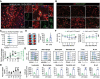






References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Research Materials
Miscellaneous
