Antimicrobial resistance and genomic investigation of non-typhoidal Salmonella isolated from outpatients in Shaoxing city, China
- PMID: 36176509
- PMCID: PMC9513250
- DOI: 10.3389/fpubh.2022.988317
Antimicrobial resistance and genomic investigation of non-typhoidal Salmonella isolated from outpatients in Shaoxing city, China
Abstract
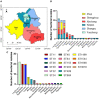
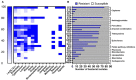
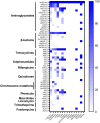
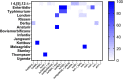
Human non-typhoidal salmonellosis is among the leading cause of morbidity and mortality worldwide, resulting in huge economic losses and threatening the public health systems. To date, epidemiological characteristics of non-typhoidal Salmonella (NTS) implicated in human salmonellosis in China are still obscure. Herein, we investigate the antimicrobial resistance and genomic features of NTS isolated from outpatients in Shaoxing city in 2020. Eighty-seven Salmonella isolates were recovered and tested against 28 different antimicrobial agents, representing 12 categories. The results showed high resistance to cefazolin (86.21%), streptomycin (81.61%), ampicillin (77.01%), ampicillin-sulbactam (74.71%), doxycycline (72.41%), tetracycline (71.26%), and levofloxacin (70.11%). Moreover, 83.91% of isolates were resistant to ≥3 categories, which were considered multi-drug resistant (MDR). Whole-genome sequencing (WGS) combined with bioinformatic analysis was used to predict serovars, MLST types, plasmid replicons, antimicrobial resistance genes, and virulence genes, in addition to the construction of phylogenomic to determine the epidemiological relatedness between isolates. Fifteen serovars and 16 STs were identified, with the dominance of S. I 4, [5], 12:i:- ST34 (25.29%), S. Enteritidis ST11 (22.99%), and S. Typhimurium ST19. Additionally, 50 resistance genes representing ten categories were detected with a high prevalence of aac(6')-Iaa (100%), bla TEM-1B (65.52%), and tet(A) (52.87%), encoding resistance to aminoglycosides, β-lactams, and tetracyclines, respectively; in addition to chromosomic mutations affecting gyrA gene. Moreover, we showed the detection of 18 different plasmids with the dominance of IncFIB(S) and IncFII(S) (39.08%). Interestingly, all isolates harbor the typical virulence genes implicated in the virulence mechanisms of Salmonella, while one isolate of S. Jangwani contains the cdtB gene encoding typhoid toxin production. Furthermore, the phylogenomic analysis showed that all isolates of the same serovar are very close to each other and clustered together in the same clade. Together, we showed a high incidence of MDR among the studied isolates which is alarming for public health services and is a major threat to the currently available treatments to deal with human salmonellosis; hence, efforts should be gathered to further introduce WGS in routinely monitoring of AMR Salmonella in the medical field in order to enhance the effectiveness of surveillance systems and to limit the spread of MDR clones.
Keywords: antimicrobial resistance; gastroenteritis; non-typhoidal Salmonella; public health; salmonellosis; virulence; whole genome sequencing.
Copyright © 2022 Chen, Ed-Dra, Zhou, Wu, Zhang and Yue.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures

References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical

