The Cys-N-degron pathway modulates pexophagy through the N-terminal oxidation and arginylation of ACAD10
- PMID: 36184612
- PMCID: PMC10262816
- DOI: 10.1080/15548627.2022.2126617
The Cys-N-degron pathway modulates pexophagy through the N-terminal oxidation and arginylation of ACAD10
Abstract
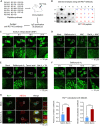
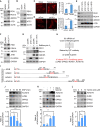
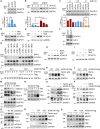
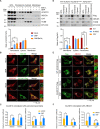
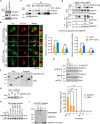
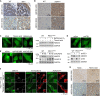
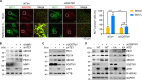
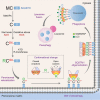
In the N-degron pathway, N-recognins recognize cognate substrates for degradation via the ubiquitin (Ub)-proteasome system (UPS) or the autophagy-lysosome system (hereafter autophagy). We have recently shown that the autophagy receptor SQSTM1/p62 (sequestosome 1) is an N-recognin that binds the N-terminal arginine (Nt-Arg) as an N-degron to modulate autophagic proteolysis. Here, we show that the N-degron pathway mediates pexophagy, in which damaged peroxisomal fragments are degraded by autophagy under normal and oxidative stress conditions. This degradative process initiates when the Nt-Cys of ACAD10 (acyl-CoA dehydrogenase family, member 10), a receptor in pexophagy, is oxidized into Cys sulfinic (CysO2) or sulfonic acid (CysO3) by ADO (2-aminoethanethiol (cysteamine) dioxygenase). Under oxidative stress, the Nt-Cys of ACAD10 is chemically oxidized by reactive oxygen species (ROS). The oxidized Nt-Cys2 is arginylated by ATE1-encoded R-transferases, generating the RCOX N-degron. RCOX-ACAD10 marks the site of pexophagy via the interaction with PEX5 and binds the ZZ domain of SQSTM1/p62, recruiting LC3+-autophagic membranes. In mice, knockout of either Ate1 responsible for Nt-arginylation or Sqstm1/p62 leads to increased levels of peroxisomes. In the cells from patients with peroxisome biogenesis disorders (PBDs), characterized by peroxisomal loss due to uncontrolled pexophagy, inhibition of either ATE1 or SQSTM1/p62 was sufficient to recover the level of peroxisomes. Our results demonstrate that the Cys-N-degron pathway generates an N-degron that regulates the removal of damaged peroxisomal membranes along with their contents. We suggest that tannic acid, a commercially available drug on the market, has a potential to treat PBDs through its activity to inhibit ATE1 R-transferases.Abbreviations: ACAA1, acetyl-Coenzyme A acyltransferase 1; ACAD, acyl-Coenzyme A dehydrogenase; ADO, 2-aminoethanethiol (cysteamine) dioxygenase; ATE1, arginyltransferase 1; CDO1, cysteine dioxygenase type 1; ER, endoplasmic reticulum; LIR, LC3-interacting region; MOXD1, monooxygenase, DBH-like 1; NAC, N-acetyl-cysteine; Nt-Arg, N-terminal arginine; Nt-Cys, N-terminal cysteine; PB1, Phox and Bem1p; PBD, peroxisome biogenesis disorder; PCO, plant cysteine oxidase; PDI, protein disulfide isomerase; PTS, peroxisomal targeting signal; R-COX, Nt-Arg-CysOX; RNS, reactive nitrogen species; ROS, reactive oxygen species; SNP, sodium nitroprusside; UBA, ubiquitin-associated; UPS, ubiquitinproteasome system.
Keywords: Acyl-CoA dehydrogenase family, member 10; N-degron pathway; oxidative stress; peroxisomal biogenesis disorders; peroxisome; pexophagy.
Conflict of interest statement
No potential conflict of interest was reported by the author(s).
Figures

References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
Miscellaneous
