BRCA1-associated RING domain-1 (BARD1) loss and GBP1 expression enhance sensitivity to DNA damage in Ewing sarcoma
- PMID: 36187937
- PMCID: PMC9524505
- DOI: 10.1158/2767-9764.crc-21-0047
BRCA1-associated RING domain-1 (BARD1) loss and GBP1 expression enhance sensitivity to DNA damage in Ewing sarcoma
Abstract
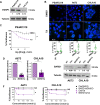
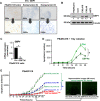
Ewing sarcoma is a fusion oncoprotein-driven primary bone tumor. A subset of patients (~10%) with Ewing sarcoma are known to harbor germline variants in a growing number of genes involved in DNA damage repair. We recently reported our discovery of a germline mutation in the DNA damage repair protein BARD1 (BRCA1-associated RING domain-1) in a patient with Ewing sarcoma. BARD1 is recruited to the site of DNA double stranded breaks via the poly(ADP-ribose) polymerase (PARP) protein and plays a critical role in DNA damage response pathways including homologous recombination. We thus questioned the impact of BARD1 loss on Ewing cell sensitivity to DNA damage and the Ewing sarcoma transcriptome. We demonstrate that PSaRC318 cells, a novel patient-derived cell line harboring a pathogenic BARD1 variant, are sensitive to PARP inhibition and by testing the effect of BARD1 depletion in additional Ewing sarcoma cell lines, we confirm that BARD1 loss enhances cell sensitivity to PARP inhibition plus radiation. Additionally, RNA-seq analysis revealed that loss of BARD1 results in the upregulation of GBP1 (guanylate-binding protein 1), a protein whose expression is associated with variable response to therapy depending on the adult carcinoma subtype examined. Here, we demonstrate that GBP1 contributes to the enhanced sensitivity of BARD1 deficient Ewing cells to DNA damage. Together, our findings demonstrate the impact of loss-of function mutations in DNA damage repair genes, such as BARD1, on Ewing sarcoma treatment response.
Keywords: BARD1; DNA damage; Ewing sarcoma; GBP1; PARPi.
Conflict of interest statement
Conflict of Interest Statement: Consulting for Ipsen and Bayer (KAJ) and Honoraria from Takeda and Foundation Medicine (KAJ). Remaining authors report no conflicts of interest to disclose.
Figures



References
-
- Grünewald TGP, Cidre-Aranaz F, Surdez D, Tomazou EM, De Álava E, Kovar H, et al. . Ewing sarcoma. Nat Rev Dis Primers 2018;4:5. - PubMed
-
- Grier HE, Krailo MD, Tarbell NJ, Link MP, Fryer CJH, Pritchard DJ, et al. . Addition of ifosfamide and etoposide to standard chemotherapy for Ewing's sarcoma and primitive neuroectodermal tumor of bone. N Engl J Med 2003;348:694–701. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases
Research Materials
Miscellaneous

