S6K1 phosphorylates Cdk1 and MSH6 to regulate DNA repair
- PMID: 36189922
- PMCID: PMC9529248
- DOI: 10.7554/eLife.79128
S6K1 phosphorylates Cdk1 and MSH6 to regulate DNA repair
Abstract
The mTORC1 substrate, S6 Kinase 1 (S6K1), is involved in the regulation of cell growth, ribosome biogenesis, glucose homeostasis, and adipogenesis. Accumulating evidence has suggested a role for mTORC1 signaling in the DNA damage response. This is mostly based on the findings that mTORC1 inhibitors sensitized cells to DNA damage. However, a direct role of the mTORC1-S6K1 signaling pathway in DNA repair and the mechanism by which this signaling pathway regulates DNA repair is unknown. In this study, we discovered a novel role for S6K1 in regulating DNA repair through the coordinated regulation of the cell cycle, homologous recombination (HR) DNA repair (HRR) and mismatch DNA repair (MMR) mechanisms. Here, we show that S6K1 orchestrates DNA repair by phosphorylation of Cdk1 at serine 39, causing G2/M cell cycle arrest enabling homologous recombination and by phosphorylation of MSH6 at serine 309, enhancing MMR. Moreover, breast cancer cells harboring RPS6KB1 gene amplification show increased resistance to several DNA damaging agents and S6K1 expression is associated with poor survival of breast cancer patients treated with chemotherapy. Our findings reveal an unexpected function of S6K1 in the DNA repair pathway, serving as a tumorigenic barrier by safeguarding genomic stability.
Keywords: CDK1; DNA repair; MSH2; MSH6; S6K1; biochemistry; cancer biology; cdk1(cdc2); chemical biology; dna repair; mTORC1; msh2; msh6; s6k1.
Plain language summary
Damage to the DNA in our cells can cause harmful changes that, if unchecked, can lead to the development of cancer. To help prevent this, cellular mechanisms are in place to repair defects in the DNA. A particular process, known as the mTORC1-S6K1 pathway is suspected to be important for repair because when this pathway is blocked, cells become more sensitive to DNA damage. It is still unknown how the various proteins involved in the mTORC1-S6K1 pathway contribute to repairing DNA. One of these proteins, S6K1, is an enzyme involved in coordinating cell growth and survival. The tumor cells in some forms of breast cancer produce more of this protein than normal, suggesting that S6K1 benefits these cells’ survival. However, it is unclear exactly how the enzyme does this. Amar-Schwartz, Ben-Hur, Jbara et al. studied the role of S6K1 using genetically manipulated mouse cells and human cancer cells. These experiments showed that the protein interacts with two other proteins involved in DNA repair and activates them, regulating two different repair mechanisms and protecting cells against damage. These results might explain why some breast cancer tumors are resistant to radiotherapy and chemotherapy treatments, which aim to kill tumor cells by damaging their DNA. If this is the case, these findings could help clinicians choose more effective treatment options for people with cancers that produce additional S6K1. In the future, drugs that block the activity of the enzyme could make cancer cells more susceptible to chemotherapy.
© 2022, Amar-Schwartz, Ben Hur, Jbara et al.
Conflict of interest statement
AA, VB, AJ, YC, GB, EA, ZS, BM, FH, AS, MA, MB, RW, RA, TG, RK No competing interests declared
Figures
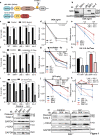
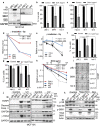
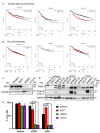
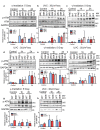
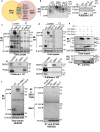



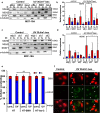


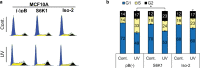
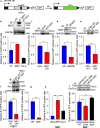
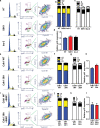

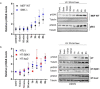
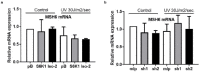
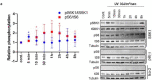
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Research Materials
Miscellaneous

