Species-informative SNP markers for characterising freshwater prawns of genus Macrobrachium in Cameroon
- PMID: 36190939
- PMCID: PMC9529149
- DOI: 10.1371/journal.pone.0263540
Species-informative SNP markers for characterising freshwater prawns of genus Macrobrachium in Cameroon
Abstract
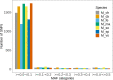
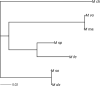
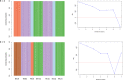
Single Nucleotide Polymorphisms (SNPs) are now popular for a myriad of applications in animal and plant species including, ancestry assignment, conservation genetics, breeding, and traceability of animal products. The objective of this study was to develop a customized cost-effective SNP panel for genetic characterisation of Macrobrachium species in Cameroon. The SNPs identified in a previous characterization study were screened as viable candidates for the reduced panel. Starting from a full set of 1,814 SNPs, a total of 72 core SNPs were chosen using conventional approaches: allele frequency differentials, minor allele frequency profiles, and Wright's Fst statistics. The discriminatory power of reduced set of informative SNPs were then tested using the admixture analysis, principal component analysis, and discriminant analysis of principal components. The panel of prioritised SNP markers (i.e., N = 72 SNPs) distinguished Macrobrachium species with 100% accuracy. However, large sample size is needed to identify more informative SNPs for discriminating genetically closely related species, including M. macrobrachion versus M. vollenhovenii and M. sollaudii versus M. dux. Overall, the findings in this study show that we can accurately characterise Macrobrachium using a small set of core SNPs which could be useful for this economically important species in Cameroon. Given the results obtained in this study, a larger independent validation sample set will be needed to confirm the discriminative capacity of this SNP panel for wider commercial and research applications.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures





References
-
- HOLTHUIS, L. B. 1980. FAO species catalogue. Volume 1-Shrimps and prawns of the world. An annotated catalogue of species of interest to fisheries.
-
- MARCH J. G., PRINGLE C. M., TOWNSEND M. J. & WILSON A. I. 2002. Effects of freshwater shrimp assemblages on benthic communities along an altitudinal gradient of a tropical island stream. Freshwater Biology, 47, 377–390.
-
- WOWOR D., MUTHU V., MEIER R., BALKE M., CAI Y. & NG P. K. 2009. Evolution of life history traits in Asian freshwater prawns of the genus Macrobrachium (Crustacea: Decapoda: Palaemonidae) based on multilocus molecular phylogenetic analysis. Molecular phylogenetics and evolution, 52, 340–350. doi: 10.1016/j.ympev.2009.01.002 - DOI - PubMed
-
- DE GRAVE S. & FRANSEN C. 2011. Carideorum catalogus: the recent species of the dendrobranchiate, stenopodidean, procarididean and caridean shrimps (Crustacea: Decapoda), NCB Naturalis Leiden.
-
- SAUNG M. H. H., KUNJURAMANVIJAYAMMA J. & LAY K. K. 2021. Two new species of Macrobrachium Spence Bate (Decapoda: Palaemonidae) from Ayeyarwady River, Myanmar with a note on Macrobrachium lamarrei (H. Milne Edwards 1837). Zoologischer Anzeiger, 293, 112–123.
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Miscellaneous

