iSnoDi-LSGT: identifying snoRNA-disease associations based on local similarity constraints and global topological constraints
- PMID: 36192132
- PMCID: PMC9670808
- DOI: 10.1261/rna.079325.122
iSnoDi-LSGT: identifying snoRNA-disease associations based on local similarity constraints and global topological constraints
Abstract
Growing evidence proves that small nucleolar RNAs (snoRNAs) have important functions in various biological processes, the malfunction of which leads to the emergence and development of complex diseases. However, identifying snoRNA-disease associations is an ongoing challenging task due to the considerable time- and money-consuming biological experiments. Therefore, it is urgent to design efficient and economical methods for the identification of snoRNA-disease associations. In this regard, we propose a computational method named iSnoDi-LSGT, which utilizes snoRNA sequence similarity and disease similarity as local similarity constraints. The iSnoDi-LSGT predictor further employs network embedding technology to extract topological features of snoRNAs and diseases, based on which snoRNA topological similarity and disease topological similarity are calculated as global topological constraints. To the best of our knowledge, the iSnoDi-LSGT is the first computational method for snoRNA-disease association identification. The experimental results indicate that the iSnoDi-LSGT predictor can effectively predict unknown snoRNA-disease associations. The web server of the iSnoDi-LSGT predictor is freely available at http://bliulab.net/iSnoDi-LSGT.
Keywords: global topological constraint; local similarity constraint; network embedding technology; snoRNA-disease association identification.
© 2022 Zhang and Liu; Published by Cold Spring Harbor Laboratory Press for the RNA Society.
Figures
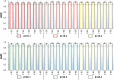


References
-
- Bouchard-Bourelle P, Desjardins-Henri C, Mathurin-St-Pierre D, Deschamps-Francoeur G, Fafard-Couture É, Garant J-M, Elela SA, Scott MS. 2020. snoDB: an interactive database of human snoRNA sequences, abundance and interactions. Nucleic Acids Res 48: D220–D225. 10.1093/nar/gkz884 - DOI - PMC - PubMed
-
- Bragantini B, Charron C, Bourguet M, Paul A, Tiotiu D, Rothé B, Marty H, Terral G, Hessmann S, Decourty L, et al. 2021. The box C/D snoRNP assembly factor Bcd1 interacts with the histone chaperone Rtt106 and controls its transcription dependent activity. Nat Commun 12: 1859. 10.1038/s41467-021-22077-4 - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
