The Drosophila ZAD zinc finger protein Kipferl guides Rhino to piRNA clusters
- PMID: 36193674
- PMCID: PMC9531945
- DOI: 10.7554/eLife.80067
The Drosophila ZAD zinc finger protein Kipferl guides Rhino to piRNA clusters
Abstract
RNA interference systems depend on the synthesis of small RNA precursors whose sequences define the target spectrum of these silencing pathways. The Drosophila Heterochromatin Protein 1 (HP1) variant Rhino permits transcription of PIWI-interacting RNA (piRNA) precursors within transposon-rich heterochromatic loci in germline cells. Current models propose that Rhino's specific chromatin occupancy at piRNA source loci is determined by histone marks and maternally inherited piRNAs, but also imply the existence of other, undiscovered specificity cues. Here, we identify a member of the diverse family of zinc finger associated domain (ZAD)-C2H2 zinc finger proteins, Kipferl, as critical Rhino cofactor in ovaries. By binding to guanosine-rich DNA motifs and interacting with the Rhino chromodomain, Kipferl recruits Rhino to specific loci and stabilizes it on chromatin. In kipferl mutant flies, Rhino is lost from most of its target chromatin loci and instead accumulates on pericentromeric Satellite arrays, resulting in decreased levels of transposon targeting piRNAs and impaired fertility. Our findings reveal that DNA sequence, in addition to the H3K9me3 mark, determines the identity of piRNA source loci and provide insight into how Rhino might be caught in the crossfire of genetic conflicts.
Keywords: D. melanogaster; HP1 proteins; ZAD zinc finger proteins; chromosomes; gene expression; genetics; genomics; germline biology; heterochromatin formation; piRNA pathway; transposon biology; zinc finger proteins.
Plain language summary
The genes within our DNA encode the essentials of our body plan and how each task in the body is achieved. However, our genome also contains many repetitive regions of DNA that do not encode functional genes. Some of these regions are genetic parasites known as transposons that try to multiply and spread around the DNA of their host. To prevent transposon DNA from interfering with the way the body operates, humans and other animals have evolved elaborate defense mechanisms to identify transposons and prevent them from multiplying. In one such mechanism, known as the piRNA pathway, the host makes small molecules known as piRNAs that have sequences complementary to those of transposons, and act as guides to silence the transposons. The instructions to make these piRNAs are stored in the form of transposon fragments in dedicated regions of host DNA called piRNA clusters. These clusters thereby act as genetic memory, allowing the host to recognize and silence specific transposons in other locations within the host’s genome. In fruit flies, a protein called Rhino binds to piRNA clusters that are densely packed to allow piRNAs to be made. However, it remained unclear how Rhino is able to identify and bind to piRNA clusters, but not to other similarly densely packed regions of DNA. Baumgartner et al. used a combination of genetic, genomic, and imaging approaches to study how Rhino finds its way in the fruit fly genome. They found that another protein called Kipferl interacts with Rhino and is required for Rhino to bind to nearly all piRNA clusters. Since Kipferl can by itself bind to the sequences that Rhino needs to find, the results suggest that Kipferl acts to recruit and initiate Rhino binding within densely packed piRNA clusters. Further experiments found that, in flies lacking Kipferl, Rhino binds to regions of DNA called Satellite repeats, hinting that these selfish sequences may compete for Rhino for their own benefit. The finding that Kipferl and Rhino work together to define the memory system of the piRNA pathway strongly advances our understanding of how a sequence-specific defense system based on small RNAs can be established.
© 2022, Baumgartner et al.
Conflict of interest statement
LB, DH, SP, CY, PD, JB No competing interests declared
Figures


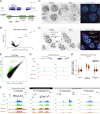


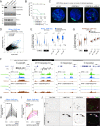
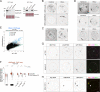




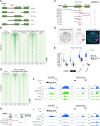
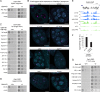


Comment in
-
More than just an inert dense region.Elife. 2022 Oct 14;11:e83076. doi: 10.7554/eLife.83076. Elife. 2022. PMID: 36239700 Free PMC article.
References
-
- Abad JP, Agudo M, Molina I, Losada A, Ripoll P, Villasante A. Pericentromeric regions containing 1.688 satellite DNA sequences show anti-kinetochore antibody staining in prometaphase chromosomes of Drosophila melanogaster. Molecular & General Genetics. 2000;264:371–377. doi: 10.1007/s004380000331. - DOI - PubMed
-
- Akkouche A, Mugat B, Barckmann B, Varela-Chavez C, Li B, Raffel R, Pélisson A, Chambeyron S. Piwi is required during Drosophila embryogenesis to license dual-strand pirna clusters for transposon repression in adult ovaries. Molecular Cell. 2017;66:411–419. doi: 10.1016/j.molcel.2017.03.017. - DOI - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials

