CovInter: interaction data between coronavirus RNAs and host proteins
- PMID: 36200814
- PMCID: PMC9825556
- DOI: 10.1093/nar/gkac834
CovInter: interaction data between coronavirus RNAs and host proteins
Abstract
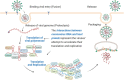
Coronavirus has brought about three massive outbreaks in the past two decades. Each step of its life cycle invariably depends on the interactions among virus and host molecules. The interaction between virus RNA and host protein (IVRHP) is unique compared to other virus-host molecular interactions and represents not only an attempt by viruses to promote their translation/replication, but also the host's endeavor to combat viral pathogenicity. In other words, there is an urgent need to develop a database for providing such IVRHP data. In this study, a new database was therefore constructed to describe the interactions between coronavirus RNAs and host proteins (CovInter). This database is unique in (a) unambiguously characterizing the interactions between virus RNA and host protein, (b) comprehensively providing experimentally validated biological function for hundreds of host proteins key in viral infection and (c) systematically quantifying the differential expression patterns (before and after infection) of these key proteins. Given the devastating and persistent threat of coronaviruses, CovInter is highly expected to fill the gap in the whole process of the 'molecular arms race' between viruses and their hosts, which will then aid in the discovery of new antiviral therapies. It's now free and publicly accessible at: https://idrblab.org/covinter/.
© The Author(s) 2022. Published by Oxford University Press on behalf of Nucleic Acids Research.
Figures

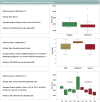
Similar articles
-
RVvictor: Virus RNA-directed molecular interactions for RNA virus infection.Comput Biol Med. 2024 Feb;169:107886. doi: 10.1016/j.compbiomed.2023.107886. Epub 2023 Dec 20. Comput Biol Med. 2024. PMID: 38157777
-
The current landscape of coronavirus-host protein-protein interactions.J Transl Med. 2020 Aug 18;18(1):319. doi: 10.1186/s12967-020-02480-z. J Transl Med. 2020. PMID: 32811513 Free PMC article. Review.
-
Determination of host proteins composing the microenvironment of coronavirus replicase complexes by proximity-labeling.Elife. 2019 Jan 11;8:e42037. doi: 10.7554/eLife.42037. Elife. 2019. PMID: 30632963 Free PMC article.
-
Host Factors in Coronavirus Replication.Curr Top Microbiol Immunol. 2018;419:1-42. doi: 10.1007/82_2017_25. Curr Top Microbiol Immunol. 2018. PMID: 28643204 Free PMC article. Review.
-
Continuous and Discontinuous RNA Synthesis in Coronaviruses.Annu Rev Virol. 2015 Nov;2(1):265-88. doi: 10.1146/annurev-virology-100114-055218. Annu Rev Virol. 2015. PMID: 26958916 Free PMC article. Review.
Cited by
-
CovEpiAb: a comprehensive database and analysis resource for immune epitopes and antibodies of human coronaviruses.Brief Bioinform. 2024 Mar 27;25(3):bbae183. doi: 10.1093/bib/bbae183. Brief Bioinform. 2024. PMID: 38653491 Free PMC article.
-
SYNBIP 2.0: epitopes mapping, sequence expansion and scaffolds discovery for synthetic binding protein innovation.Nucleic Acids Res. 2025 Jan 6;53(D1):D595-D603. doi: 10.1093/nar/gkae893. Nucleic Acids Res. 2025. PMID: 39413165 Free PMC article.
-
Alpha and gamma mangostins inhibit wild-type B SARS-CoV-2 more effectively than the SARS-CoV-2 variants and the major target is unlikely the 3C-like protease.Heliyon. 2024 May 27;10(11):e31987. doi: 10.1016/j.heliyon.2024.e31987. eCollection 2024 Jun 15. Heliyon. 2024. PMID: 38867992 Free PMC article.
-
The 2023 Nucleic Acids Research Database Issue and the online molecular biology database collection.Nucleic Acids Res. 2023 Jan 6;51(D1):D1-D8. doi: 10.1093/nar/gkac1186. Nucleic Acids Res. 2023. PMID: 36624667 Free PMC article.
-
Entropy driven cooperativity effect in multi-site drug optimization targeting SARS-CoV-2 papain-like protease.Cell Mol Life Sci. 2023 Oct 5;80(11):313. doi: 10.1007/s00018-023-04985-4. Cell Mol Life Sci. 2023. PMID: 37796323 Free PMC article.
References
-
- Oudshoorn D., Rijs K., Limpens R., Groen K., Koster A.J., Snijder E.J., Kikkert M., Barcena M.. Expression and cleavage of middle east respiratory syndrome coronavirus nsp3-4 polyprotein induce the formation of double-membrane vesicles that mimic those associated with coronaviral RNA replication. Mbio. 2017; 8:e01658-17. - PMC - PubMed
-
- Rodriguez-Morales A.J., Bonilla-Aldana D.K., Balbin-Ramon G.J., Rabaan A.A., Sah R., Paniz-Mondolfi A., Pagliano P., Esposito S.. History is repeating itself: probable zoonotic spillover as the cause of the 2019 novel coronavirus epidemic. Infez. Med. 2020; 28:3–5. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Miscellaneous

