ChemPert: mapping between chemical perturbation and transcriptional response for non-cancer cells
- PMID: 36200827
- PMCID: PMC9825489
- DOI: 10.1093/nar/gkac862
ChemPert: mapping between chemical perturbation and transcriptional response for non-cancer cells
Abstract
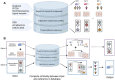
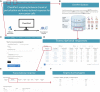
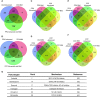
Prior knowledge of perturbation data can significantly assist in inferring the relationship between chemical perturbations and their specific transcriptional response. However, current databases mostly contain cancer cell lines, which are unsuitable for the aforementioned inference in non-cancer cells, such as cells related to non-cancer disease, immunology and aging. Here, we present ChemPert (https://chempert.uni.lu/), a database consisting of 82 270 transcriptional signatures in response to 2566 unique perturbagens (drugs, small molecules and protein ligands) across 167 non-cancer cell types, as well as the protein targets of 57 818 perturbagens. In addition, we develop a computational tool that leverages the non-cancer cell datasets, which enables more accurate predictions of perturbation responses and drugs in non-cancer cells compared to those based onto cancer databases. In particular, ChemPert correctly predicted drug effects for treating hepatitis and novel drugs for osteoarthritis. The ChemPert web interface is user-friendly and allows easy access of the entire datasets and the computational tool, providing valuable resources for both experimental researchers who wish to find datasets relevant to their research and computational researchers who need comprehensive non-cancer perturbation transcriptomics datasets for developing novel algorithms. Overall, ChemPert will facilitate future in silico compound screening for non-cancer cells.
© The Author(s) 2022. Published by Oxford University Press on behalf of Nucleic Acids Research.
Figures

Similar articles
-
LINPS: a database for cancer-cell-specific perturbations of biological networks.Database (Oxford). 2021 Aug 20;2021:baab048. doi: 10.1093/database/baab048. Database (Oxford). 2021. PMID: 34415996 Free PMC article.
-
PerturBase: a comprehensive database for single-cell perturbation data analysis and visualization.Nucleic Acids Res. 2025 Jan 6;53(D1):D1099-D1111. doi: 10.1093/nar/gkae858. Nucleic Acids Res. 2025. PMID: 39377396 Free PMC article.
-
Predicting transcriptional responses to novel chemical perturbations using deep generative model for drug discovery.Nat Commun. 2024 Oct 26;15(1):9256. doi: 10.1038/s41467-024-53457-1. Nat Commun. 2024. PMID: 39462106 Free PMC article.
-
Free resources to assist structure-based virtual ligand screening experiments.Curr Protein Pept Sci. 2007 Aug;8(4):381-411. doi: 10.2174/138920307781369391. Curr Protein Pept Sci. 2007. PMID: 17696871 Review.
-
Software packages for quantitative microarray-based gene expression analysis.Curr Pharm Biotechnol. 2003 Dec;4(6):417-37. doi: 10.2174/1389201033377436. Curr Pharm Biotechnol. 2003. PMID: 14683435 Review.
Cited by
-
PC3T: a signature-driven predictor of chemical compounds for cellular transition.Commun Biol. 2023 Sep 27;6(1):989. doi: 10.1038/s42003-023-05225-y. Commun Biol. 2023. PMID: 37758874 Free PMC article.
-
Deep representation learning of chemical-induced transcriptional profile for phenotype-based drug discovery.Nat Commun. 2024 Jun 25;15(1):5378. doi: 10.1038/s41467-024-49620-3. Nat Commun. 2024. PMID: 38918369 Free PMC article.
-
Reversal gene expression assessment for drug repurposing, a case study of glioblastoma.J Transl Med. 2025 Jan 7;23(1):25. doi: 10.1186/s12967-024-06046-1. J Transl Med. 2025. PMID: 39773231 Free PMC article.
-
On the correspondence between the transcriptomic response of a compound and its effects on its targets.BMC Bioinformatics. 2023 May 19;24(1):207. doi: 10.1186/s12859-023-05337-6. BMC Bioinformatics. 2023. PMID: 37208587 Free PMC article.
-
Towards an interpretable deep learning model of cancer.NPJ Precis Oncol. 2025 Feb 14;9(1):46. doi: 10.1038/s41698-025-00822-y. NPJ Precis Oncol. 2025. PMID: 39948231 Free PMC article. Review.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

