Tissue clearing of human iPSC-derived organ-chips enables high resolution imaging and analysis
- PMID: 36205191
- PMCID: PMC9595176
- DOI: 10.1039/d2lc00116k
Tissue clearing of human iPSC-derived organ-chips enables high resolution imaging and analysis
Abstract
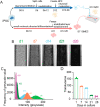
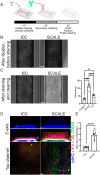
Engineered microfluidic organ-chips enable increased cellular diversity and function of human stem cell-derived tissues grown in vitro. These three dimensional (3D) cultures, however, are met with unique challenges in visualization and quantification of cellular proteins. Due to the dense 3D nature of cultured nervous tissue, classical methods of immunocytochemistry are complicated by sub-optimal light and antibody penetrance as well as image acquisition parameters. In addition, complex polydimethylsiloxane scaffolding surrounding the tissue of interest can prohibit high resolution microscopy and spatial analysis. Hyperhydration tissue clearing methods have been developed to mitigate similar challenges of in vivo tissue imaging. Here, we describe an adaptation of this approach to efficiently clear human pluripotent stem cell-derived neural tissues grown on organ-chips. We also describe critical imaging considerations when designing signal intensity-based approaches to complex 3D architectures inherent in organ-chips. To determine morphological and anatomical features of cells grown in organ-chips, we have developed a reliable protocol for chip sectioning and high-resolution microscopic acquisition and analysis.
Conflict of interest statement
Cedars-Sinai owns a minority stock interest in Emulate, the company that produces the study's microfluidic organ-chips. An officer of Cedars-Sinai also serves on Emulate's Board of Directors. Emulate provided no financial support for this research. Cedars-Sinai and Emulate have patents filed related to this work.
Figures

References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

