Multi-omics analysis of the oncogenic value of copper Metabolism-Related protein COMMD2 in human cancers
- PMID: 36205192
- PMCID: PMC10242316
- DOI: 10.1002/cam4.5320
Multi-omics analysis of the oncogenic value of copper Metabolism-Related protein COMMD2 in human cancers
Abstract
Background: The copper metabolism MURR1 domain (COMMD) protein family is involved in tumorigenicity of malignant tumors. However, as the member of COMMD, the role of COMMD2 in human tumors remains unknown.
Methods: We used The Cancer Genome Atlas (TCGA), Genotype Tissue Expression (GTEx), Human Protein Atlas (HPA) database, Cancer Cell Line Encyclopedia (CCLE) platform, univariate Cox regression analysis, Kaplan-Meier curve, cBioPortal, UALCAN database, Sangerbox online platform, GSCA database gene set enrichment analysis (GSEA), and GeneMANIA to analyze the expression of COMMD2, its prognostic values, genomic alteration patterns, and the correlation with tumor stemness, tumor mutational burden (TMB), microsatellite instability (MSI), and immune infiltrates, drug sensitivity, and gene function enrichment in pan-cancer. qRT-PCR, CCK-8, EdU, wound healing, and transwell migration assays were performed to confirm the function of COMMD2.
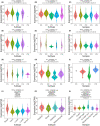
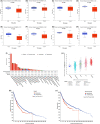
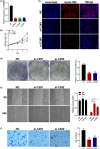
Results: COMMD2 was strongly expressed in most cancer types. Elevated COMMD2 expression affects the prognosis, clinicopathological stage, and molecular or immune subtypes of various tumors. Moreover, promoter hypomethylation and mutations in the COMMD2 gene may be associated with its high expression and poor survival. Additionally, we discovered that COMMD2 expression was linked to tumor stemness, TMB, MSI, immune cell infiltration, immune-checkpoint inhibitors, and drug sensitivity in pan-cancer. Furthermore, the COMMD2 gene co-expression network is constructed with GSEA analysis, displaying significant interaction of COMMD2 with E2F targets, G2-M checkpoint, and mitotic spindle in bladder cancer (BLCA). Finally, RNA interference data showed suppression of COMMD2 prevented proliferation and migration of BLCA and uterine corpus endometrial carcinoma (UCEC) cells.
Conclusion: Our findings shed light on the COMMD2 functions in human cancers and demonstrate that it is a promising prognostic biomarker and therapeutic target in pan-cancer.
Keywords: COMMD2; bioinformatics; immune infiltration; pan-cancer; prognostic biomarker.
© 2022 The Authors. Cancer Medicine published by John Wiley & Sons Ltd.
Conflict of interest statement
The authors declared no conflict of interest.
Figures








References
-
- Bray F, Ferlay J, Soerjomataram I, Siegel RL, Torre LA, Jemal A. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2018;68(6):394‐424. - PubMed
-
- Blum A, Wang P, Zenklusen JC. SnapShot: TCGA‐analyzed tumors. Cell. 2018;173(2):530. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases

