Acoustic force spectroscopy reveals subtle differences in cellulose unbinding behavior of carbohydrate-binding modules
- PMID: 36215467
- PMCID: PMC9586272
- DOI: 10.1073/pnas.2117467119
Acoustic force spectroscopy reveals subtle differences in cellulose unbinding behavior of carbohydrate-binding modules
Abstract
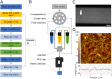
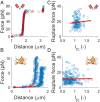
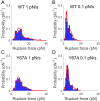
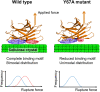
Protein adsorption to solid carbohydrate interfaces is critical to many biological processes, particularly in biomass deconstruction. To engineer more-efficient enzymes for biomass deconstruction into sugars, it is necessary to characterize the complex protein-carbohydrate interfacial interactions. A carbohydrate-binding module (CBM) is often associated with microbial surface-tethered cellulosomes or secreted cellulase enzymes to enhance substrate accessibility. However, it is not well known how CBMs recognize, bind, and dissociate from polysaccharides to facilitate efficient cellulolytic activity, due to the lack of mechanistic understanding and a suitable toolkit to study CBM-substrate interactions. Our work outlines a general approach to study the unbinding behavior of CBMs from polysaccharide surfaces using a highly multiplexed single-molecule force spectroscopy assay. Here, we apply acoustic force spectroscopy (AFS) to probe a Clostridium thermocellum cellulosomal scaffoldin protein (CBM3a) and measure its dissociation from nanocellulose surfaces at physiologically relevant, low force loading rates. An automated microfluidic setup and method for uniform deposition of insoluble polysaccharides on the AFS chip surfaces are demonstrated. The rupture forces of wild-type CBM3a, and its Y67A mutant, unbinding from nanocellulose surfaces suggests distinct multimodal CBM binding conformations, with structural mechanisms further explored using molecular dynamics simulations. Applying classical dynamic force spectroscopy theory, the single-molecule unbinding rate at zero force is extrapolated and found to agree with bulk equilibrium unbinding rates estimated independently using quartz crystal microbalance with dissipation monitoring. However, our results also highlight critical limitations of applying classical theory to explain the highly multivalent binding interactions for cellulose-CBM bond rupture forces exceeding 15 pN.
Keywords: acoustic force spectroscopy; biofuels; carbohydrate-binding module; nanocellulose; single-molecule force spectroscopy.
Conflict of interest statement
The authors declare no competing interest.
Figures



Similar articles
-
Measurements of single molecular affinity interactions between carbohydrate-binding modules and crystalline cellulose fibrils.Phys Chem Chem Phys. 2013 May 7;15(17):6508-15. doi: 10.1039/c3cp51072g. Phys Chem Chem Phys. 2013. PMID: 23532050
-
Imaging and measuring single-molecule interaction between a carbohydrate-binding module and natural plant cell wall cellulose.J Phys Chem B. 2012 Aug 23;116(33):9949-56. doi: 10.1021/jp304686q. Epub 2012 Aug 10. J Phys Chem B. 2012. PMID: 22849362
-
Molecular origins of reduced activity and binding commitment of processive cellulases and associated carbohydrate-binding proteins to cellulose III.J Biol Chem. 2021 Jan-Jun;296:100431. doi: 10.1016/j.jbc.2021.100431. Epub 2021 Feb 19. J Biol Chem. 2021. PMID: 33610545 Free PMC article.
-
Carbohydrate-binding modules facilitate the enzymatic hydrolysis of lignocellulosic biomass: Releasing reducing sugars and dissociative lignin available for producing biofuels and chemicals.Biotechnol Adv. 2023 Jul-Aug;65:108126. doi: 10.1016/j.biotechadv.2023.108126. Epub 2023 Mar 14. Biotechnol Adv. 2023. PMID: 36921877 Review.
-
Carbohydrate-binding modules of fungal cellulases: occurrence in nature, function, and relevance in industrial biomass conversion.Adv Appl Microbiol. 2014;88:103-65. doi: 10.1016/B978-0-12-800260-5.00004-8. Adv Appl Microbiol. 2014. PMID: 24767427 Review.
Cited by
-
Combining DNA scaffolds and acoustic force spectroscopy to characterize individual protein bonds.Biophys J. 2023 Jun 20;122(12):2518-2530. doi: 10.1016/j.bpj.2023.05.004. Epub 2023 Jun 7. Biophys J. 2023. PMID: 37290437 Free PMC article.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

