BNIP3L/NIX regulates both mitophagy and pexophagy
- PMID: 36215693
- PMCID: PMC9753467
- DOI: 10.15252/embj.2022111115
BNIP3L/NIX regulates both mitophagy and pexophagy
Abstract
Mitochondria and peroxisomes are closely related metabolic organelles, both in terms of origin and in terms of function. Mitochondria and peroxisomes can also be turned over by autophagy, in processes termed mitophagy and pexophagy, respectively. However, despite their close relationship, it is not known if both organelles are turned over under similar conditions, and if so, how this might be coordinated molecularly. Here, we find that multiple selective autophagy pathways are activated upon iron chelation and show that mitophagy and pexophagy occur in a BNIP3L/NIX-dependent manner. We reveal that the outer mitochondrial membrane-anchored NIX protein, previously described as a mitophagy receptor, also independently localises to peroxisomes and drives pexophagy. We show this process happens in vivo, with mouse tissue that lacks NIX having a higher peroxisomal content. We further show that pexophagy is stimulated under the same physiological conditions that activate mitophagy, including cardiomyocyte and erythrocyte differentiation. Taken together, our work uncovers a dual role for NIX, not only in mitophagy but also in pexophagy, thus illustrating the interconnection between selective autophagy pathways.
Keywords: autophagy; mitochondria; mitophagy; peroxisomes; pexophagy.
© 2022 The Authors. Published under the terms of the CC BY 4.0 license.
Figures

- A
Representative immunoblots and quantifications (below) of the indicated proteins in lysates of ARPE‐19 cells treated with 1 mM DFP for the indicated time (h, hours).
- B, C
Representative confocal images of ARPE‐19 cells stably expressing the mito‐QC reporter (B), or the pexo‐QC reporter (C) treated with 1 mM DFP for the indicated time (h). Enlarged images of the area outlined in white are shown in the lower corners. The “red‐only” mask is generated during images analysis and highlights red‐only puncta (autolysosomes) based on the mCherry/GFP ratio and thresholding criteria. Nuclei were stained in blue (Hoechst). Scale bar: 10 μm. At right, quantification of total red‐only punctate per cell (mitolysosomes or pexolysosomes).
- D
ARPE‐19 cells were transfected with 50 pmol of non‐targeting siRNA (siNT) or 50 pmol of siRNA targeting ATG13 or ULK1. 48 h post knockdown, cells were treated with 1 mM DFP for an additional 48 h. Cell lysates were subject to immunoblotting with the indicated antibodies. Quantification is shown on the right.

- A
Representative immunoblots of the indicated proteins in lysates of ARPE‐19 cells treated 48 h with 1 mM DFP.
- B, C
Representative immunoblots of the indicated proteins in lysates of HeLa and SH‐SY5Y cells (B) or ARPE‐19 cells (C), treated with 1 mM DFP or 50 μM Phenentroline (Phen) for the indicated time (h).
- D
Representative immunoblots of the indicated proteins in lysates of ARPE‐19 cells stimulated with hypoxia (0.8%) for 3 days.
- E
Representative confocal images of ARPE‐19 cells stably expressing the pexo‐QC reporter stimulated with hypoxia (0.8%) for 3 days and at right, quantification of total red‐only punctate per cell (pexolysosomes). Enlarged images of the area outlined in white are shown in the lower corners.
- F, G
Representative confocal images of ARPE‐19 cells stably expressing the pexo‐QC reporter treated with 1 mM DFP for 48 h and immunostained with peroxisomal markers (F, Catalase and PMP70), or lysosomal marker (G, LAMP1). Enlarged images of the area outlined in white are shown on the right or below.
- H, I
ARPE‐19 cells expressing the pexo‐QC reporter were transfected with 50 pmol of non‐targeting siRNA (siNT) or 50 pmol of siRNA targeting ATG13 or ULK1. 48 h post transfection, cells were treated with 1 mM DFP for an additional 48 h and analysed by immunoblot (H) or by flow cytometry (I). Representative dot plots are shown after analysing GFP and mCherry signals. The percentage of cell underdoing pexophagy (purple population) is indicated in bold purple on each dot plot.

- A
Representative immunoblots of the indicated proteins in lysates of WT or HIF1α KO U2OS cells, treated with 1 mM DFP for 48 h.
- B
Representative confocal images of WT or HIF1α KO U2OS cells stably expressing the pexo‐QC reporter and treated with 1 mM DFP for 48 h (left panel) and flow cytometry analysis of the mCherry/GFP ratio (right panel).
- C
Representative immunoblots (left panel) and quantification (right panel) of the indicated proteins in lysates of WT or two NIX KO ARPE‐19 clones (Cl), treated with 1 mM DFP for the indicated time (h).
- D, E
Representative confocal images of WT or NIX KO ARPE‐19 cells stably expressing the mito‐QC reporter (D) or the pexo‐QC reporter (E) treated with 1 mM DFP for 48 h (left panel) and flow cytometry analysis of the mCherry/GFP ratio (right panel).
- F
Representative immunoblots (left panel) and quantification (right panel) of the indicated proteins. NIX KO ARPE‐19 cells (Cl 31) stably expressing a pBabe empty vector (∅) or a pBabe NIX vector, were treated with 1 mM DFP for 48 h prior lysis. Two biological replicates are shown in the immunoblot.

- A
Representative immunoblots of the indicated proteins in lysates of ARPE‐19 cells transfected with 50 pmol of non‐targeting siRNA (NT) or 50 pmol of siRNA targeting NIX or BNIP3. 48 h post transfection, cells were treated with 1 mM DFP for an additional 48 h.
- B, C
Quantification by flow cytometry of the mCherry/GFP ratio in ARPE‐19 cells expressing the mito‐QC reporter (B) or the pexo‐QC reporter (C) and treated as in (A).
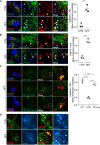
- A, B
Representative confocal images of untreated (CTRL) and DFP‐treated ARPE‐19 cells (1 mM, 48 h), stably expressing GFP‐WT NIX (green) and stained with anti‐catalase antibody (A, red) or anti‐PMP70 antibody (B, red). At right, quantification of Pearson correlation coefficients between NIX and the peroxisomal markers catalase and PMP70, with or without DFP treatment for 48 h. Peroxisomes enriched for GFP‐NIX or having minimal signal for GFP‐NIX are depicted with white arrowheads or white arrows, respectively.
- C
Representative confocal images of ARPE‐19 cells stably expressing GFP‐WT NIX, GFP‐NIXΔTM or GFP‐NIX TM only (green), treated with DFP (1 mM) for 48 h and immunostained with anti‐catalase antibody (red). At right, quantification of Pearson correlation coefficients between the green and the red signal. Overall data are mean ± s.d.; Statistical significance is displayed as **P ≤ 0.01; ***P ≤ 0.001; ****P ≤ 0.0001; ns, not significant. n = 3 biological replicates with > 15 cells per replicate, (A, B) unpaired t‐test, two tailed; (C) one‐way ANOVA, Tukey's multiple comparisons test.
- D
Representative confocal images of untreated (CTRL) and DFP‐treated ARPE‐19 cells (1 mM, 48 h), stably expressing GFP‐ WT NIX (green) and immunostained with a mitochondrial marker (ATPB, cyan) and peroxisomal marker (catalase, red).
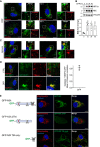
- A
Representative confocal images of ARPE‐19 cells immunostained with an anti‐NIX antibody (green) and anti‐catalase antibody (red) after 1 mM DFP for the indicated time (h). Peroxisomes enriched for NIX or having minimal signal for NIX are depicted with yellow or white arrowheads, respectively.
- B
Representative immunoblots and quantifications (below) of the indicated proteins in lysates of cells treated as (A). Data are mean ± s.d.; n = 4 biological replicates. *P ≤ 0.05; **P ≤ 0.01; ns, not significant (one‐way ANOVA, Tukey's multiple comparisons test).
- C
Representative confocal images of ARPE‐19 cells WT (right panel) or stably expressing GFP‐NIX (left panel) and stimulated with hypoxia for 3 days. Peroxisomes were stained with anti‐catalase antibody (red) and endogenous NIX were stained with anti‐NIX antibody (green, right panel).
- D
Representative confocal images of untreated (CTRL) and DFP‐treated ARPE‐19 cells (1 mM, 48 h), stably expressing GFP‐BNIP3 (green) and stained with anti‐catalase antibody (red). At right, quantification of Pearson correlation coefficients between BNIP3 and the peroxisomal marker catalase after DFP treatment for 48 h.
- E
Schematic representation of the NIX mutants. Right panel, confocal images of ARPE‐19 cells stably expressing GFP‐NIX mutants and immunostained with a mitochondrial marker (ATPB, red).
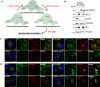
- A
Schematic illustration of mitochondria depletion in cells overexpressing Parkin (PRKN). Cells are pre‐treated for 24 h with CCCP (20 μM), and after 1 day of recovery (CCCP washout), treated for 48 h with 1 mM DFP.
- B
Representative immunoblots of the indicated proteins in lysates of WT or HA‐Parkin (PRKN)‐ARPE‐19 cells treated as in (A).
- C, D
Representative confocal images of HA‐Parkin (PRKN) ARPE‐19 cells treated as in (A) and stained with anti‐NIX antibody (green) and anti‐ATPB antibody (red, C) or anti‐Catalase antibody (red, D). Enlarged images of the area outlined in white are shown on the right. Nuclei were stained in blue (Hoechst) and scale bars: 10 μm.
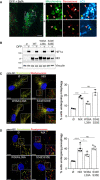
- A
Representative confocal images of ARPE‐19 cells stably expressing mCherry‐SKL (peroxisomes, red), treated with 1 mM DFP for 48 h and 50 nM Bafilomycin A (BafA) for the last 6 h prior imaging. Mitochondria were stained with anti‐COXIV antibody (green) and autophagosomes with anti‐LC3 antibody (cyan). Green‐only, red‐only, and green and red positive structures are depicted with yellow, red and white arrowheads, respectively. Scale bars for (i) and (ii) is 2 μm. Enlarged images of the area outlined in white are shown on the right.
- B
Representative immunoblots of the indicated proteins in lysates of NIX KO ARPE‐19 cells (Cl 31) stably expressing a pBabe Flag vector (∅), a pBabe Flag‐NIX vector (NIX), a pBabe Flag‐NIX W36A\L39A vector (W36A\L39A) or a pBabe Flag‐NIX S34E\S35E vector (S34E\S35E), and treated with 1 mM DFP for 48 h prior lysis.
- C, D
Representative confocal images of cells as in (B), stably expressing the mito‐QC reporter (C) or the pexo‐QC reporter (D) and treated with 1 mM DFP for 24 h (C) or 48 h (D) (left panel) and flow cytometry analysis of the mCherry/GFP ratio (right panel). Enlarged images of the area outlined in white are shown in the lower corners. Nuclei were stained in blue (Hoechst) and scale bars: 10 μm.

- A
Representative immunoblots and quantification of the indicated proteins in lysates of ARPE‐19 cells treated with 1 mM DFP, 20 μM CCCP, 10 μM Antimycin and 5 μM Oligomycin A (O/A), 15 μM Ivermectin for 24 h.
- B
Representative confocal images of ARPE‐19 cells treated as in A and stained with anti‐Ubiquitin antibody (FK2).
- C
Representative confocal images of ARPE‐19 cells stably expressing the pexo‐QC reporter and treated with 1 mM DFP or 10 μM Antimycin and 5 μM Oligomycin A (O/A) for 48 h. Below, flow cytometry analysis of the mCherry/GFP ratio.
- D
Representative confocal images of ARPE‐19 cells treated with EBSS for 24 h or 1 mM DFP for 48 h and immunostained with anti‐catalase (red) and anti‐p62 (green) antibodies. Autophagosomes containing peroxisomes are depicted with white arrows while autophagosomes not colocalizing with peroxisome marker are pointed with white arrowheads. Enlarged images of the area outlined in white are shown on the right.

- A
Representative confocal images of optical section from WT and NIX KO mouse retina immunostained with peroxisomal markers (PMP70 and catalase) or a lysosomal marker LAMP1. Nuclei are coloured in white (DAPI). RPE, retinal pigment epithelium; ONL, outer nuclear layer; OPL, outer plexiform layer; INL; inner nuclear layer; IPL, inner plexiform layer; RGC, retinal ganglion cells. Scale bar: 20 mm.
- B
Quantification of signal area in the ONL and INL from images as (A). Each data point represents a single mouse (n = 3–4 per group).
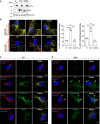
- A
Representative immunoblots of the indicated proteins in lysates of NIX KO ARPE‐19 cells (Cl 31) stably expressing a pBabe Flag vector (∅), a pBabe Flag‐NIX vector (NIX), a pBabe Flag‐NIX G204A vector (G204A), and treated with 1 mM DFP for 48 h prior lysis.
- B
Representative confocal images of cells as in (A), stably expressing the mito‐QC reporter or the pexo‐QC reporter and treated with 1 mM DFP for 48 h (left panel) and flow cytometry analysis of the mCherry/GFP ratio (right panel). Enlarged images of the area outlined in white are shown in the lower corners.
- C, D
Representative confocal images of NIX KO ARPE‐19 cells (Cl 31) stably expressing a pBabe Flag vector (∅), a pBabe Flag‐NIX vector (NIX), a pBabe Flag‐NIX W36A\L39A vector (W36A\L39A) or a pBabe Flag‐NIX G204A vector (G204A), untreated (C) or treated with 1 mM DFP for 48 h (D) and immunostained with a mitochondrial marker (C, ATPB in red) or a peroxisomal marker (D, Catalase in red) and with an anti‐NIX antibody (green).
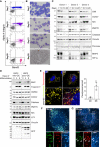
- A
Representative flow cytometry data of in vitro human erythroid cells cultured for 7, 10 and 13 days, as well as of reticulocytes obtained after 20 days in culture. The dot plots show unstained cells (black), cells stained with IgG control antibodies (grey) and cells stained with antibodies against alpha 4 integrin and Band 3 (coloured).
- B
Representative images of erythroblasts or reticulocytes as in (A), prepared by cytospin and stained with May‐Grünwald's stain and Giemsa's stain. Scale bars: 10 μm.
- C
Immunoblots of the indicated proteins in lysates of erythroblasts or reticulocytes obtained from three different donors.
- D
Representative immunoblots of the indicated proteins in lysates of mito‐QC‐ or pexo‐QC‐H9c2 cells during differentiation.
- E
Representative confocal images of pexo‐QC reporter H9c2 cells before (D0) and after 8 days of differentiation (D8). Nuclei were stained in blue (Hoechst). Red‐only puncta (pexolysosomes) are depicted with white arrowheads. At right, quantification of total number of pexolysosomes counted per cell analysed during cadiomyocyte differentiation. Overall data are mean ± s.d.; n = 3 biological replicates with > 85 cells per condition and replicate, unpaired t‐test, two‐tailed. **P ≤ 0.01.
- F, G
Representative confocal images of differentiated pexo‐QC reporter H9c2 cells immunostained with anti‐catalase (F) or anti‐LAMP1 (G). White arrowheads indicate structures positive for both red‐puncta and LAMP1. Enlarged images of the area outlined in white are shown below.
References
-
- Aviram I, Shaklai N (1981) The association of human erythrocyte catalase with the cell membrane. Arch Biochem Biophys 212: 329–337 - PubMed
-
- Bingol B, Tea JS, Phu L, Reichelt M, Bakalarski CE, Song Q, Foreman O, Kirkpatrick DS, Sheng M (2014) The mitochondrial deubiquitinase USP30 opposes Parkin‐mediated mitophagy. Nature 510: 370–375 - PubMed
-
- Braschi E, Goyon V, Zunino R, Mohanty A, Xu L, McBride HM (2010) Vps35 mediates vesicle transport between the mitochondria and peroxisomes. Curr Biol 20: 1310–1315 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

