Genomic and Epidemiological Features of Two Dominant Methicillin-Susceptible Staphylococcus aureus Clones from a Neonatal Intensive Care Unit Surveillance Effort
- PMID: 36218345
- PMCID: PMC9769867
- DOI: 10.1128/msphere.00409-22
Genomic and Epidemiological Features of Two Dominant Methicillin-Susceptible Staphylococcus aureus Clones from a Neonatal Intensive Care Unit Surveillance Effort
Abstract
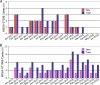
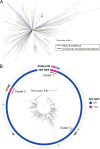
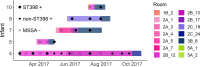
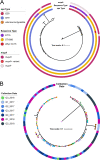
Methicillin-susceptible Staphylococcus aureus (MSSA) is a more prevalent neonatal intensive care unit (NICU) pathogen than methicillin-resistant S. aureus (MRSA). However, the introduction and spread of MSSA, the role of systematic decolonization, and optimal infection prevention and control strategies remain incompletely understood. We previously screened infants hospitalized in a university-affiliated level III to IV NICU twice monthly over 18 months for S. aureus colonization and identified several prevalent staphylococcal protein A (spa) types. Here, we performed whole-genome sequencing (WGS) and phylogenetic comparisons of 140 isolates from predominant spa types t279, t1451, and t571 to examine possible transmission routes and identify genomic and epidemiologic features associated with the spread of dominant clones. We identified two major MSSA clones: sequence type 398 (ST398), common in the local community, and ST1898, not previously encountered in the region. ST398 NICU isolates formed distinct clusters with closely related community isolates from previously published data sets, suggesting multiple sources of acquisition, such as family members or staff, including residents of the local community. In contrast, ST1898 isolates were nearly identical, pointing to clonal expansion within the NICU. Almost all ST1898 isolates harbored plasmids encoding mupirocin resistance (mupA), suggesting an association between the proliferation of this clone and decolonization efforts with mupirocin. Comparative genomics indicated genotype-specific pathways of introduction and spread of MSSA via community-associated (ST398) or health care-associated (ST1898) sources and the potential role of mupirocin resistance in dissemination of ST1898. Future surveillance efforts could benefit from routine genotyping to inform clone-specific infection prevention strategies. IMPORTANCE Methicillin-susceptible Staphylococcus aureus (MSSA) is a significant pathogen in neonates. However, surveillance efforts in neonatal intensive care units (NICUs) have focused primarily on methicillin-resistant S. aureus (MRSA), limiting our understanding of colonizing and infectious MSSA clones which are prevalent in the NICU. Here, we identify two dominant colonizing MSSA clones during an 18-month surveillance effort in a level III to IV NICU, ST398 and ST1898. Using genomic surveillance and phylogenetic analysis, coupled with epidemiological investigation, we found that these two sequence types had distinct modes of spread, namely the suggested exchange with community reservoirs for ST398 and the contribution of antibiotic resistance to dissemination of ST1898 in the health care setting. This study highlights the additional benefits of whole-genome surveillance for colonizing pathogens, beyond routine species identification and genotyping, to inform targeted infection prevention strategies.
Keywords: MSSA; NICU; genomic surveillance; methicillin-susceptible Staphylococcus aureus; mupirocin resistance; neonatal intensive care unit (NICU).
Conflict of interest statement
The authors declare no conflict of interest.
Figures
Similar articles
-
Spa Typing of Staphylococcus aureus in a Neonatal Intensive Care Unit During Routine Surveillance.J Pediatric Infect Dis Soc. 2021 Aug 17;10(7):766-773. doi: 10.1093/jpids/piab014. J Pediatric Infect Dis Soc. 2021. PMID: 34129043 Free PMC article.
-
Comparative Genomics of Borderline Oxacillin-Resistant Staphylococcus aureus Detected during a Pseudo-outbreak of Methicillin-Resistant S. aureus in a Neonatal Intensive Care Unit.mBio. 2022 Feb 22;13(1):e0319621. doi: 10.1128/mbio.03196-21. Epub 2022 Jan 18. mBio. 2022. PMID: 35038924 Free PMC article.
-
Emergence of community-associated methicillin-resistant Staphylococcus aureus strains in the neonatal intensive care unit: an infection prevention and patient safety challenge.Clin Microbiol Infect. 2016 Jul;22(7):645.e1-8. doi: 10.1016/j.cmi.2016.04.013. Epub 2016 Apr 25. Clin Microbiol Infect. 2016. PMID: 27126609 Free PMC article.
-
Mupirocin resistance in Staphylococcus aureus: A systematic review and meta-analysis.J Glob Antimicrob Resist. 2020 Mar;20:238-247. doi: 10.1016/j.jgar.2019.07.032. Epub 2019 Aug 20. J Glob Antimicrob Resist. 2020. PMID: 31442624
-
Microbiome dysbiosis: a modifiable state and target to prevent Staphylococcus aureus infections and other diseases in neonates.J Perinatol. 2024 Jan;44(1):125-130. doi: 10.1038/s41372-023-01810-5. Epub 2023 Oct 30. J Perinatol. 2024. PMID: 37904005 Free PMC article. Review.
Cited by
-
Genomic and Phenotypic Characterization of Mupirocin-resistant Staphylococcus aureus Clinical Isolates.Open Forum Infect Dis. 2025 Jul 15;12(7):ofaf374. doi: 10.1093/ofid/ofaf374. eCollection 2025 Jul. Open Forum Infect Dis. 2025. PMID: 40698033 Free PMC article.
-
Whole-genome comparison of two same-genotype macrolide-resistant Bordetella pertussis isolates collected in Japan.PLoS One. 2024 Feb 15;19(2):e0298147. doi: 10.1371/journal.pone.0298147. eCollection 2024. PLoS One. 2024. PMID: 38359004 Free PMC article.
-
Factors associated with persistent blood stream infection in the Neonatal Intensive Care Unit.Res Sq [Preprint]. 2025 Jul 1:rs.3.rs-6946297. doi: 10.21203/rs.3.rs-6946297/v1. Res Sq. 2025. PMID: 40630520 Free PMC article. Preprint.
-
The mutational landscape of Staphylococcus aureus during colonisation.Nat Commun. 2025 Jan 13;16(1):302. doi: 10.1038/s41467-024-55186-x. Nat Commun. 2025. PMID: 39805814 Free PMC article.
References
-
- McMullan BJ, Bowen A, Blyth CC, Van Hal S, Korman TM, Buttery J, Voss L, Roberts S, Cooper C, Tong SYC, Turnidge J. 2016. Epidemiology and mortality of Staphylococcus aureus bacteremia in Australian and New Zealand children. JAMA Pediatr 170:979–986. doi:10.1001/jamapediatrics.2016.1477. - DOI - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
