Dynamic in Situ Confinement Triggers Ligand-Free Neuropeptide Receptor Signaling
- PMID: 36219818
- PMCID: PMC9614963
- DOI: 10.1021/acs.nanolett.2c03506
Dynamic in Situ Confinement Triggers Ligand-Free Neuropeptide Receptor Signaling
Abstract
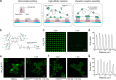
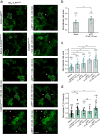
Membrane receptor clustering is fundamental to cell-cell communication; however, the physiological function of receptor clustering in cell signaling remains enigmatic. Here, we developed a dynamic platform to induce cluster formation of neuropeptide Y2 hormone receptors (Y2R) in situ by a chelator nanotool. The multivalent interaction enabled a dynamic exchange of histidine-tagged Y2R within the clusters. Fast Y2R enrichment in clustered areas triggered ligand-independent signaling as determined by an increase in cytosolic calcium and cell migration. Notably, the calcium and motility response to ligand-induced activation was amplified in preclustered cells, suggesting a key role of receptor clustering in sensitizing the dose response to lower ligand concentrations. Ligand-independent versus ligand-induced signaling differed in the binding of arrestin-3 as a downstream effector, which was recruited to the clusters only in the presence of the ligand. This approach allows in situ receptor clustering, raising the possibility to explore different receptor activation modalities.
Keywords: G protein-coupled receptors; membrane organization; phase separation; receptor condensates; receptor dynamics.
Conflict of interest statement
The authors declare no competing financial interest.
Figures



References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

