Genomic characterization of two community-acquired methicillin-resistant Staphylococcus aureus with novel sequence types in Kenya
- PMID: 36226152
- PMCID: PMC9548584
- DOI: 10.3389/fmed.2022.966283
Genomic characterization of two community-acquired methicillin-resistant Staphylococcus aureus with novel sequence types in Kenya
Abstract
Staphylococcus aureus is a clinically important bacteria with high antimicrobial resistance (AMR) challenge globally. The emergence of methicillin-resistant Staphylococcus aureus (MRSA) clones with unique sequence types have been identified in the community showing evidence that the epidemiology of MRSA globally is changing and requires continual surveillance. We utilized whole genome sequencing to characterize two community acquired-MRSA (CA-MRSA) strains isolated from wound swabs from community-onset infections in two health facilities in Kenya. The two strains belonged to multilocus sequence type (MLST) sequence type (ST) 7460, and ST 7635. The resistance genes detected showed that the novel STs are carriers of clinically relevant resistance genes. Linezolid and mupirocin resistance was observed, yet mupirocin is not commonly used in the country. Mutations within resistance genes were also detected and the pathogenicity toward the human host matched various pathogenic global S. aureus families, e.g., S. aureus subsp. aureus USA300. Multidrug efflux transporters, important in antimicrobial resistance including restriction enzymes type I and type IV were detected. Plasmids identified showed similarities with the plasmids in other clinically significant non-staphylococcal species, such as Pseudomonas aeruginosa, Escherichia coli, Morganella morganii, and Enterococcus faecium. Both STs belong to clonal complex 8 (CC8) which is the most successful MRSA clone in Kenya. Spa type t30 to which ST 7635 belongs has not been reported in the country. The results of this study further highlight the need for epidemiological studies to reveal circulating strains and antimicrobial resistance spread between hospitals and the community. The genomic research highlights resistance to anti-staphylococcal broad-spectrum antimicrobials not used frequently in the country, jeopardizing successful MRSA treatment since most health facilities do not perform genotypic resistance tests for routine patient management. Preliminary insights into unidentified STs of CA-MRSA in Kenya show the need for molecular epidemiological surveillance studies to further understand the diversity of S. aureus in Africa.
Keywords: AMR; CA-MRSA; Kenya; WGS; novel sequence.
Copyright © 2022 Njenga, Nyasinga, Munshi, Muraya, Omuse, Ngugi and Revathi.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures
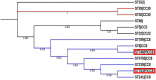
References
-
- Becker REN, Bubeck Wardenburg J. Staphylococcus aureus and the skin: a longstanding and complex interaction. Skinmed. (2015) 13:111–9. - PubMed
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

