Epigenetic Modulation of Opioid Receptors by Drugs of Abuse
- PMID: 36233105
- PMCID: PMC9569510
- DOI: 10.3390/ijms231911804
Epigenetic Modulation of Opioid Receptors by Drugs of Abuse
Abstract
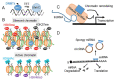
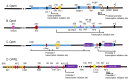
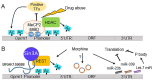
Chronic exposure to drugs of abuse produces profound changes in gene expression and neural activity associated with drug-seeking and taking behavior. Dysregulation of opioid receptor gene expression is commonly observed across a variety of abused substances including opioids, cocaine, and alcohol. Early studies in cultured cells showed that the spatial and temporal gene expression of opioid receptors are regulated by epigenetic mechanisms including DNA and histone modifications and non-coding RNAs. Accumulating evidence indicate that drugs of abuse can modulate opioid receptor gene expression by targeting various epigenetic regulatory networks. Based on current cellular and animal models of substance use disorder and clinical evidence, this review summarizes how chronic drug exposure alters the gene expression of mu, delta, kappa, and nociceptin receptors via DNA and histone modifications. The influence of drugs of abuse on epigenetic modulators, such as non-coding RNAs and transcription factors, is also presented. Finally, the therapeutic potential of manipulating epigenetic processes as an avenue to treat substance use disorder is discussed.
Keywords: DNA methylation; drugs of abuse; epigenetics; gene expression; histone modifications; noncoding RNAs; opioid receptors.
Conflict of interest statement
The authors declare no conflict of interest.
Figures
Similar articles
-
Dynorphin/KOP and nociceptin/NOP gene expression and epigenetic changes by cocaine in rat striatum and nucleus accumbens.Prog Neuropsychopharmacol Biol Psychiatry. 2014 Mar 3;49:36-46. doi: 10.1016/j.pnpbp.2013.10.016. Epub 2013 Nov 1. Prog Neuropsychopharmacol Biol Psychiatry. 2014. PMID: 24184686
-
15 years of genetic approaches in vivo for addiction research: Opioid receptor and peptide gene knockout in mouse models of drug abuse.Neuropharmacology. 2014 Jan;76 Pt B(0 0):204-17. doi: 10.1016/j.neuropharm.2013.08.028. Epub 2013 Sep 10. Neuropharmacology. 2014. PMID: 24035914 Free PMC article. Review.
-
Epigenetic Mechanisms of Opioid Addiction.Biol Psychiatry. 2020 Jan 1;87(1):22-33. doi: 10.1016/j.biopsych.2019.06.027. Epub 2019 Jul 8. Biol Psychiatry. 2020. PMID: 31477236 Free PMC article. Review.
-
Epigenetics of drug abuse: predisposition or response.Pharmacogenomics. 2012 Jul;13(10):1149-60. doi: 10.2217/pgs.12.94. Pharmacogenomics. 2012. PMID: 22909205 Free PMC article. Review.
-
Analysis of epigenetic mechanisms regulating opioid receptor gene transcription.Methods Mol Biol. 2015;1230:39-51. doi: 10.1007/978-1-4939-1708-2_3. Methods Mol Biol. 2015. PMID: 25293314
Cited by
-
Oxycodone, an opioid like the others?Front Psychiatry. 2023 Dec 13;14:1229439. doi: 10.3389/fpsyt.2023.1229439. eCollection 2023. Front Psychiatry. 2023. PMID: 38152360 Free PMC article. Review.
-
The Impact of Opioids on Epigenetic Modulation in Myocardial Ischemia and Reperfusion Injury: Focus on Non-coding RNAs.J Cardiovasc Transl Res. 2025 Apr 8. doi: 10.1007/s12265-025-10609-y. Online ahead of print. J Cardiovasc Transl Res. 2025. PMID: 40198537 Review.
-
Unraveling the Hippocampal Molecular and Cellular Alterations behind Tramadol and Tapentadol Neurobehavioral Toxicity.Pharmaceuticals (Basel). 2024 Jun 17;17(6):796. doi: 10.3390/ph17060796. Pharmaceuticals (Basel). 2024. PMID: 38931463 Free PMC article.
-
Opioids and opioid receptors; understanding pharmacological mechanisms as a key to therapeutic advances and mitigation of the misuse crisis.BJA Open. 2023 May 17;6:100141. doi: 10.1016/j.bjao.2023.100141. eCollection 2023 Jun. BJA Open. 2023. PMID: 37588171 Free PMC article. Review.
-
Disrupted neuregulin 1-ErbB4 signaling: Consequences of prenatal morphine exposure in rat pups and molecular gateway to neurological impairment.Toxicol Rep. 2024 Jul 8;13:101687. doi: 10.1016/j.toxrep.2024.101687. eCollection 2024 Dec. Toxicol Rep. 2024. PMID: 39109071 Free PMC article.
References
-
- Fattore L., Fadda P., Antinori S., Fratta W. Role of opioid receptors in the reinstatement of opioid-seeking behavior: An overview. Methods Mol. Biol. 2015;1230:281–293. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Research Materials

