Evasion of neutralizing antibody responses by the SARS-CoV-2 BA.2.75 variant
- PMID: 36240764
- PMCID: PMC9515334
- DOI: 10.1016/j.chom.2022.09.015
Evasion of neutralizing antibody responses by the SARS-CoV-2 BA.2.75 variant
Abstract
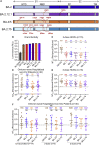
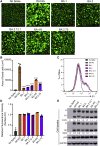
The newly emerged BA.2.75 severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) variant contains 9 additional mutations in its spike (S) protein compared to the ancestral BA.2 variant. Here, we examine the neutralizing antibody escape of BA.2.75 in mRNA-vaccinated and BA.1-infected individuals, as well as the molecular basis underlying functional changes in S. Notably, BA.2.75 exhibits enhanced neutralization resistance over BA.2 but less than the BA.4/5 variant. The G446S and N460K mutations of BA.2.75 are primarily responsible for its enhanced resistance to neutralizing antibodies. The R493Q mutation, a reversion to the prototype sequence, reduces BA.2.75 neutralization resistance. The impact of these mutations is consistent with their locations in common neutralizing antibody epitopes. Further, BA.2.75 shows enhanced cell-cell fusion over BA.2, driven largely by the N460K mutation, which enhances S processing. Structural modeling reveals enhanced receptor contacts introduced by N460K, suggesting a mechanism of potentiated receptor utilization and syncytia formation.
Keywords: BA.2.75; Omicron; SARS-CoV-2; cell-cell fusion; mRNA booster; mRNAvaccine; neutralizing antibodies.
Copyright © 2022 The Author(s). Published by Elsevier Inc. All rights reserved.
Conflict of interest statement
Declaration of interests The authors declare no competing interests.
Figures


Update of
-
Evasion of Neutralizing Antibody Response by the SARS-CoV-2 BA.2.75 Variant.bioRxiv [Preprint]. 2022 Aug 15:2022.08.14.503921. doi: 10.1101/2022.08.14.503921. bioRxiv. 2022. Update in: Cell Host Microbe. 2022 Nov 9;30(11):1518-1526.e4. doi: 10.1016/j.chom.2022.09.015. PMID: 36032970 Free PMC article. Updated. Preprint.
References
-
- Centers for Disease Control and Prevention COVID data tracker. United States Department of Health and Human Services. 2022 https://covid.cdc.gov/covid-data-tracker
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Miscellaneous

