Cryoelectron microscopy structures of a human neutralizing antibody bound to MERS-CoV spike glycoprotein
- PMID: 36246239
- PMCID: PMC9554411
- DOI: 10.3389/fmicb.2022.988298
Cryoelectron microscopy structures of a human neutralizing antibody bound to MERS-CoV spike glycoprotein
Abstract
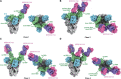
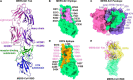
Neutralizing monoclonal antibodies (mAbs) against highly pathogenic coronaviruses represent promising candidates for clinical intervention. Here, we isolated a potent neutralizing monoclonal antibody, MERS-S41, from a yeast displayed scFv library using the S protein as a bait. To uncover the neutralization mechanism, we determined structures of MERS-S41 Fab in complex with the trimeric spike glycoprotein by cryoelectron microscopy (cryo-EM). We observed four distinct classes of the complex structure, which showed that the MERS-S41 Fab bound to the "up" receptor binding domain (RBD) with full saturation and also bound to an accessible partially lifted "down" RBD, providing a structural basis for understanding how mAbs bind to trimeric spike glycoproteins. Structure analysis of the epitope and cell surface staining assays demonstrated that virus entry is blocked predominantly by direct competition with the host receptor, dipeptidyl peptidase-4 (DPP4).
Keywords: MERS-CoV; cryo-EM structures; neutralization mechanism; neutralizing antibody; spike glycoprotein.
Copyright © 2022 Zhang, Jia, Zeng, Li, Wang, Zhou, Zhang and Wang.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures


Similar articles
-
Recombinant Receptor-Binding Domains of Multiple Middle East Respiratory Syndrome Coronaviruses (MERS-CoVs) Induce Cross-Neutralizing Antibodies against Divergent Human and Camel MERS-CoVs and Antibody Escape Mutants.J Virol. 2016 Dec 16;91(1):e01651-16. doi: 10.1128/JVI.01651-16. Print 2017 Jan 1. J Virol. 2016. PMID: 27795425 Free PMC article.
-
Potent neutralization of MERS-CoV by human neutralizing monoclonal antibodies to the viral spike glycoprotein.Sci Transl Med. 2014 Apr 30;6(234):234ra59. doi: 10.1126/scitranslmed.3008140. Epub 2014 Apr 28. Sci Transl Med. 2014. PMID: 24778414
-
Structural Definition of a Unique Neutralization Epitope on the Receptor-Binding Domain of MERS-CoV Spike Glycoprotein.Cell Rep. 2018 Jul 10;24(2):441-452. doi: 10.1016/j.celrep.2018.06.041. Cell Rep. 2018. PMID: 29996104 Free PMC article.
-
Characterization of novel monoclonal antibodies against MERS-coronavirus spike protein.Virus Res. 2020 Mar;278:197863. doi: 10.1016/j.virusres.2020.197863. Epub 2020 Jan 13. Virus Res. 2020. PMID: 31945421 Free PMC article.
-
MERS-CoV spike protein: a key target for antivirals.Expert Opin Ther Targets. 2017 Feb;21(2):131-143. doi: 10.1080/14728222.2017.1271415. Epub 2016 Dec 21. Expert Opin Ther Targets. 2017. PMID: 27936982 Free PMC article. Review.
Cited by
-
Mapping immunodominant sites on the MERS-CoV spike glycoprotein targeted by infection-elicited antibodies in humans.Cell Rep. 2024 Aug 27;43(8):114530. doi: 10.1016/j.celrep.2024.114530. Epub 2024 Jul 25. Cell Rep. 2024. PMID: 39058596 Free PMC article.
-
Mapping immunodominant sites on the MERS-CoV spike glycoprotein targeted by infection-elicited antibodies in humans.bioRxiv [Preprint]. 2024 Apr 2:2024.03.31.586409. doi: 10.1101/2024.03.31.586409. bioRxiv. 2024. Update in: Cell Rep. 2024 Aug 27;43(8):114530. doi: 10.1016/j.celrep.2024.114530. PMID: 38617298 Free PMC article. Updated. Preprint.
-
Cross-neutralization ability of anti-MERS-CoV monoclonal antibodies against a variety of merbecoviruses.Front Microbiol. 2025 Jul 16;16:1593095. doi: 10.3389/fmicb.2025.1593095. eCollection 2025. Front Microbiol. 2025. PMID: 40740328 Free PMC article.
-
Human coronavirus OC43 nanobody neutralizes virus and protects mice from infection.J Virol. 2024 Jun 13;98(6):e0053124. doi: 10.1128/jvi.00531-24. Epub 2024 May 6. J Virol. 2024. PMID: 38709106 Free PMC article.
-
Protein nanoparticle vaccines induce potent neutralizing antibody responses against MERS-CoV.Cell Rep. 2024 Dec 24;43(12):115036. doi: 10.1016/j.celrep.2024.115036. Epub 2024 Dec 6. Cell Rep. 2024. PMID: 39644492 Free PMC article.
References
LinkOut - more resources
Full Text Sources
Miscellaneous

