Ubiquitin-like processing of TUG proteins as a mechanism to regulate glucose uptake and energy metabolism in fat and muscle
- PMID: 36246906
- PMCID: PMC9556833
- DOI: 10.3389/fendo.2022.1019405
Ubiquitin-like processing of TUG proteins as a mechanism to regulate glucose uptake and energy metabolism in fat and muscle
Abstract
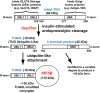
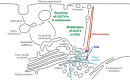
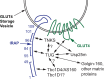
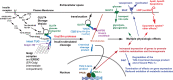
In response to insulin stimulation, fat and muscle cells mobilize GLUT4 glucose transporters to the cell surface to enhance glucose uptake. Ubiquitin-like processing of TUG (Aspscr1, UBXD9) proteins is a central mechanism to regulate this process. Here, recent advances in this area are reviewed. The data support a model in which intact TUG traps insulin-responsive "GLUT4 storage vesicles" at the Golgi matrix by binding vesicle cargoes with its N-terminus and matrix proteins with its C-terminus. Insulin stimulation liberates these vesicles by triggering endoproteolytic cleavage of TUG, mediated by the Usp25m protease. Cleavage occurs in fat and muscle cells, but not in fibroblasts or other cell types. Proteolytic processing of intact TUG generates TUGUL, a ubiquitin-like protein modifier, as the N-terminal cleavage product. In adipocytes, TUGUL modifies a single protein, the KIF5B kinesin motor, which carries GLUT4 and other vesicle cargoes to the cell surface. In muscle, this or another motor may be modified. After cleavage of intact TUG, the TUG C-terminal product is extracted from the Golgi matrix by the p97 (VCP) ATPase. In both muscle and fat, this cleavage product enters the nucleus, binds PPARγ and PGC-1α, and regulates gene expression to promote fatty acid oxidation and thermogenesis. The stability of the TUG C-terminal product is regulated by an Ate1 arginyltransferase-dependent N-degron pathway, which may create a feedback mechanism to control oxidative metabolism. Although it is now clear that TUG processing coordinates glucose uptake with other aspects of physiology and metabolism, many questions remain about how this pathway is regulated and how it is altered in metabolic disease in humans.
Keywords: GLUT4; adipocyte; glucose; insulin; membrane trafficking; muscle; p97/VCP ATPases; ubiquitin-like.
Copyright © 2022 Bogan.
Conflict of interest statement
The author declares that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures
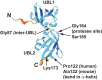
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

