Effects on Autophagy of Moxibustion at Governor Vessel Acupoints in APP/PS1double-Transgenic Alzheimer's Disease Mice through the lncRNA Six3os1/miR-511-3p/AKT3 Molecular Axis
- PMID: 36248429
- PMCID: PMC9556209
- DOI: 10.1155/2022/3881962
Effects on Autophagy of Moxibustion at Governor Vessel Acupoints in APP/PS1double-Transgenic Alzheimer's Disease Mice through the lncRNA Six3os1/miR-511-3p/AKT3 Molecular Axis
Abstract
Objective: To explore the effect and mechanism of moxibustion at acupoints of the governor vessel on lncRNA Six3os1 in amyloid precursor protein/presenilin1 (APP/PS1) double-transgenic Alzheimer's disease (AD) mice.
Methods: Twenty-four specific pathogen-free and APP/PS1 double-transgenic male mice were randomly allocated into the AD model and moxibustion groups, with 12 cases in each group. Twelve syngeneic C57BL/6J mice were selected as the control group. Mice in the moxibustion group received aconite cake-separated moxibustion at the Baihui acupoint. Suspension moxibustion was applied at Fengfu and Dazhui for 15 minutes each day. All treatments were conducted over two weeks. Control and AD model mice were routinely fed without any intervention. Behavioral observation tests were conducted before and after the intervention. The autophagosome in the hippocampus was observed using transmission electron microscopy. Immunohistochemistry was performed to detect Aβ1-42 expression. LC3B and P62 expressions were evaluated by immunofluorescence. The expression levels of the lncRNAs Six3os1, miR-511-3p, and AKT3 were detected by qRT-PCR. The differential expression of PI-3K, AKT3, mTOR, LC3B-II/I, and P62 proteins in the hippocampus was detected by western blot. The dual-luciferase assay was undertaken to examine the targeting relationships of the lncRNAs Six3os1, miR-511-3p, and AKT3.
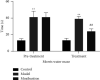
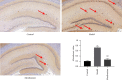
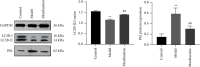
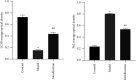
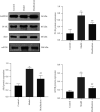
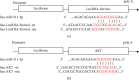
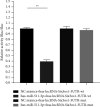
Results: Compared with the control group, the AD model showed higher escape latency in the Morris Water Maze and reduced autophagic vacuoles in the cytoplasm of hippocampal neurons (both p < 0.01). Compared with the control group, the AD model showed higher expression of Aβ1-42, the lncRNAs Six3os1, PI-3K, mTOR, P62, and AKT3 protein (all p < 0.01); but lower mir-511-3p and LC3B (both p < 0.01). Compared with the AD model group, the moxibustion group had a shorter escape latency, more autophagic bubbles in the hippocampus, and lower expression of positive Aβ1-42, the lncRNAs Six3os1, PI-3K, mTOR, P62, and AKT3 protein (all p < 0.01). In contrast, the levels of miR-511-3p and LC3B proteins were considerably increased in the moxibustion group compared to the AD model group (both p < 0.01). Based on the dual-luciferase assay, there was a targeting link among the lncRNAs Six3os1, miR-511-3p, and AKT3.
Conclusion: Moxibustion at acupoints of the governor vessel can suppress the lncRNA Six3os1 expression, promote cell autophagy, accelerate Aβ1-42 clearance and alleviate cognitive dysfunction of AD mediated by the PI3K/AKT/mTOR signaling pathway through the lncRNA Six3os1/miR-511-3p/AKT3 axis.
Copyright © 2022 Yu-Mei Jia et al.
Conflict of interest statement
The authors declare that they have no conflicts of interest.
Figures

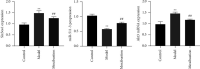
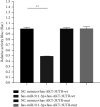
References
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

