Compensatory Genetic and Transcriptional Cytonuclear Coordination in Allopolyploid Lager Yeast (Saccharomyces pastorianus)
- PMID: 36260528
- PMCID: PMC9665066
- DOI: 10.1093/molbev/msac228
Compensatory Genetic and Transcriptional Cytonuclear Coordination in Allopolyploid Lager Yeast (Saccharomyces pastorianus)
Abstract
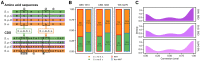
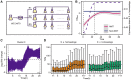
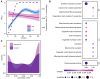
Cytonuclear coordination between biparental-nuclear genomes and uniparental-cytoplasmic organellar genomes in plants is often resolved by genetic and transcriptional cytonuclear responses. Whether this mechanism also acts in allopolyploid members of other kingdoms is not clear. Additionally, cytonuclear coordination of interleaved allopolyploid cells/individuals within the same population is underexplored. The yeast Saccharomyces pastorianus provides the opportunity to explore cytonuclear coevolution during different growth stages and from novel dimensions. Using S. pastorianus cells from multiple growth stages in the same environment, we show that nuclear mitochondria-targeted genes have undergone both asymmetric gene conversion and growth stage-specific biased expression favoring genes from the mitochondrial genome donor (Saccharomyces eubayanus). Our results suggest that cytonuclear coordination in allopolyploid lager yeast species entails an orchestrated and compensatory genetic and transcriptional evolutionary regulatory shift. The common as well as unique properties of cytonuclear coordination underlying allopolyploidy between unicellular yeasts and higher plants offers novel insights into mechanisms of cytonuclear evolution associated with allopolyploid speciation.
Keywords: allotetraploid lager yeast; cytonuclear coevolution; nuclear mitochondria-targeted gene.
© The Author(s) 2022. Published by Oxford University Press on behalf of Society for Molecular Biology and Evolution.
Figures

Similar articles
-
Industrially Applicable De Novo Lager Yeast Hybrids with a Unique Genomic Architecture: Creation and Characterization.Appl Environ Microbiol. 2021 Jan 15;87(3):e02434-20. doi: 10.1128/AEM.02434-20. Print 2021 Jan 15. Appl Environ Microbiol. 2021. PMID: 33188002 Free PMC article.
-
Cytonuclear evolution of rubisco in four allopolyploid lineages.Mol Biol Evol. 2014 Oct;31(10):2624-36. doi: 10.1093/molbev/msu207. Epub 2014 Jul 10. Mol Biol Evol. 2014. PMID: 25015644 Free PMC article.
-
Microbe Profile: Saccharomyces eubayanus, the missing link to lager beer yeasts.Microbiology (Reading). 2018 Sep;164(9):1069-1071. doi: 10.1099/mic.0.000677. Microbiology (Reading). 2018. PMID: 30175956 Free PMC article. Review.
-
Himalayan Saccharomyces eubayanus Genome Sequences Reveal Genetic Markers Explaining Heterotic Maltotriose Consumption by Saccharomyces pastorianus Hybrids.Appl Environ Microbiol. 2019 Oct 30;85(22):e01516-19. doi: 10.1128/AEM.01516-19. Print 2019 Nov 15. Appl Environ Microbiol. 2019. PMID: 31519660 Free PMC article.
-
Brewing up a storm: The genomes of lager yeasts and how they evolved.Biotechnol Adv. 2017 Jul;35(4):512-519. doi: 10.1016/j.biotechadv.2017.03.003. Epub 2017 Mar 8. Biotechnol Adv. 2017. PMID: 28284994 Review.
Cited by
-
Development of a genome-scale metabolic model for the lager hybrid yeast S. pastorianus to understand the evolution of metabolic pathways in industrial settings.mSystems. 2024 Jun 18;9(6):e0042924. doi: 10.1128/msystems.00429-24. Epub 2024 May 31. mSystems. 2024. PMID: 38819150 Free PMC article.
-
Organellar genome divergence and environmental stress induce transcriptional cytonuclear responses in wheat alloplasmic hybrids.Proc Natl Acad Sci U S A. 2025 Jun 17;122(24):e2424424122. doi: 10.1073/pnas.2424424122. Epub 2025 Jun 9. Proc Natl Acad Sci U S A. 2025. PMID: 40489605 Free PMC article.
References
-
- Adams KL, Wendel JF. 2005. Novel patterns of gene expression in polyploid plants. Trends Genet. 21:539–543. - PubMed
-
- Angelika BH, Opalińska M, Gerbeth C, Herman Josip S, Rao S, Schönfisch B, Guiard B, Schmidt O, Pfanner N, Meisinger C. 2014. Cell cycle-dependent regulation of mitochondrial preprotein translocase. Science. 346:1109–1113. - PubMed
Publication types
MeSH terms
Supplementary concepts
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

