Surveillance and Genomic Analysis of Third-Generation Cephalosporin-Resistant and Carbapenem-Resistant Klebsiella pneumoniae Complex in Germany
- PMID: 36289942
- PMCID: PMC9598256
- DOI: 10.3390/antibiotics11101286
Surveillance and Genomic Analysis of Third-Generation Cephalosporin-Resistant and Carbapenem-Resistant Klebsiella pneumoniae Complex in Germany
Abstract
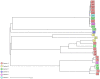
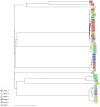
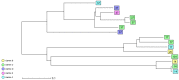
To analyse the epidemiology and population structure of third-generation cephalosporin-resistant (3GCR) and carbapenem-resistant (CR) Klebsiella pneumoniae complex isolates, patients were screened for rectal colonisation with 3GCR/CR K. pneumoniae complex on admission to six German university hospitals (2016-2019). Also collected were 3GCR/CR and susceptible K. pneumoniae isolates from patients with bloodstream infections (2016-2018). Whole-genome sequencing was performed followed by multilocus sequencing typing (MLST), core-genome MLST, and resistome and virulome analysis. The admission prevalence of 3GCR K. pneumoniae complex isolates during the 4-year study period was 0.8%, and 1.0 bloodstream infection per 1000 patient admissions was caused by K. pneumoniae complex (3GCR prevalence, 15.1%). A total of seven K. pneumoniae complex bloodstream isolates were CR (0.8%). The majority of colonising and bloodstream 3GCR isolates were identified as K. pneumoniae, 96.7% and 98.8%, respectively; the remainder were K. variicola and K. quasipneumoniae. cgMLST showed a polyclonal population of colonising and bloodstream isolates, which was also reflected by MLST and virulome analysis. CTX-M-15 was the most prevalent extended-spectrum beta-lactamase, and 29.7% of the colonising and 48.8% of the bloodstream isolates were high-risk clones. The present study provides an insight into the polyclonal 3GCR K. pneumoniae population in German hospitals.
Keywords: Klebsiella pneumoniae complex; bloodstream infections; carbapenem resistance; colonisation; third-generation cephalosporin resistance; typing.
Conflict of interest statement
C.I. has served as a consultant to MSD Sharp and Dohme and Shionogi and has received lecture fees from MSD Sharp and Dohme. S.P. consulted for IDbyDNA and received speaker’s honoraria from bioMérieux. MJGTV received research grants from 3M, Astellas Pharma, Biontech, DaVolterra, Evonik, Gilead Sciences, Glycom, Immunic, MaaT Pharma, Merck/MSD, Organobalance, Seres Therapeutics, and Takeda Pharmaceutical and speaker fees or consultation fees from Alb Fils Kliniken GmbH, Arderypharm, Astellas Pharma, Basilea, bioMérieux, DaVolterra, Farmak International Holding GmbH, Ferring, Gilead Sciences, Immunic AG, MaaT Pharma, Merck/MSD, Pfizer, Roche, Organobalance, and SocraTec R&D GmbH. The remaining authors declare no conflict of interest. The funders had no role in the design of the study; in the collection, analyses, or interpretation of data; in the writing of the manuscript; or in the decision to publish the results.
Figures

References
-
- Tacconelli E., Carrara E., Savoldi A., Harbarth S., Mendelson M., Monnet D.L., Pulcini C., Kahlmeter G., Kluytmans J., Carmeli Y., et al. Discovery, research, and development of new antibiotics: The WHO priority list of antibiotic-resistant bacteria and tuberculosis. Lancet Infect. Dis. 2018;18:318–327. doi: 10.1016/S1473-3099(17)30753-3. - DOI - PubMed
-
- Holt K.E., Wertheim H., Zadoks R.N., Baker S., Whitehouse C.A., Dance D., Jenney A., Connor T.R., Hsu L.Y., Severin J., et al. Genomic analysis of diversity, population structure, virulence, and antimicrobial resistance in Klebsiella pneumoniae, an urgent threat to public health. Proc. Natl. Acad. Sci. USA. 2015;112:E3574–E3581. doi: 10.1073/pnas.1501049112. - DOI - PMC - PubMed
Grants and funding
LinkOut - more resources
Full Text Sources

