A Novel Framework for Analysis of the Shared Genetic Background of Correlated Traits
- PMID: 36292579
- PMCID: PMC9602050
- DOI: 10.3390/genes13101694
A Novel Framework for Analysis of the Shared Genetic Background of Correlated Traits
Abstract
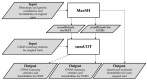
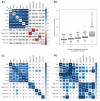
We propose a novel effective framework for the analysis of the shared genetic background for a set of genetically correlated traits using SNP-level GWAS summary statistics. This framework called SHAHER is based on the construction of a linear combination of traits by maximizing the proportion of its genetic variance explained by the shared genetic factors. SHAHER requires only full GWAS summary statistics and matrices of genetic and phenotypic correlations between traits as inputs. Our framework allows both shared and unshared genetic factors to be effectively analyzed. We tested our framework using simulation studies, compared it with previous developments, and assessed its performance using three real datasets: anthropometric traits, psychiatric conditions and lipid concentrations. SHAHER is versatile and applicable to summary statistics from GWASs with arbitrary sample sizes and sample overlaps, allows for the incorporation of different GWAS models (Cox, linear and logistic), and is computationally fast.
Keywords: GWAS; linear combination of traits; proportion of heritability explained by SGF; shared genetic component; shared heritability.
Conflict of interest statement
P.R.H.J.T. is an employee of BioAge Labs. The remaining authors declare that they have no conflict of interest.
Figures


References
-
- Tsepilov Y.A., Freidin M.B., Shadrina A.S., Sharapov S.Z., Elgaeva E.E., van Zundert J., Karssen L.C., Suri P., Williams F.M., Aulchenko Y.S. Analysis of genetically independent phenotypes identifies shared genetic factors associated with chronic musculoskeletal pain conditions. Commun. Biol. 2020;3:329. doi: 10.1038/s42003-020-1051-9. - DOI - PMC - PubMed
-
- Sampson J.N., Wheeler W.A., Yeager M., Panagiotou O., Wang Z., Berndt S.I., Lan Q., Abnet C.C., Amundadottir L.T., Figueroa J.D. Analysis of heritability and shared heritability based on genome-wide association studies for 13 cancer types. JNCI J. Natl. Cancer Inst. 2015;107:djv279. doi: 10.1093/jnci/djv279. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

