Restriction of Flaviviruses by an Interferon-Stimulated Gene SHFL/C19orf66
- PMID: 36293480
- PMCID: PMC9604422
- DOI: 10.3390/ijms232012619
Restriction of Flaviviruses by an Interferon-Stimulated Gene SHFL/C19orf66
Abstract
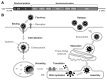
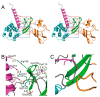
Flaviviruses (the genus Flavivirus of the Flaviviridae family) include many arthropod-borne viruses, often causing life-threatening diseases in humans, such as hemorrhaging and encephalitis. Although the flaviviruses have a significant clinical impact, it has become apparent that flavivirus replication is restricted by cellular factors induced by the interferon (IFN) response, which are called IFN-stimulated genes (ISGs). SHFL (shiftless antiviral inhibitor of ribosomal frameshifting) is a novel ISG that inhibits dengue virus (DENV), West Nile virus (WNV), Zika virus (ZIKV), and Japanese encephalitis virus (JEV) infections. Interestingly, SHFL functions as a broad-spectrum antiviral factor exhibiting suppressive activity against various types of RNA and DNA viruses. In this review, we summarize the current understanding of the molecular mechanisms by which SHFL inhibits flavivirus infection and discuss the molecular basis of the inhibitory mechanism using a predicted tertiary structure of SHFL generated by the program AlphaFold2.
Keywords: AlphaFold2; C19orf66; SHFL; flavivirus; interferon-stimulated antiviral gene.
Conflict of interest statement
The authors declare no conflict of interest.
Figures

Similar articles
-
Shiftless inhibits flavivirus replication in vitro and is neuroprotective in a mouse model of Zika virus pathogenesis.Proc Natl Acad Sci U S A. 2021 Dec 7;118(49):e2111266118. doi: 10.1073/pnas.2111266118. Proc Natl Acad Sci U S A. 2021. PMID: 34873063 Free PMC article.
-
Viperin Restricts Zika Virus and Tick-Borne Encephalitis Virus Replication by Targeting NS3 for Proteasomal Degradation.J Virol. 2018 Mar 14;92(7):e02054-17. doi: 10.1128/JVI.02054-17. Print 2018 Apr 1. J Virol. 2018. PMID: 29321318 Free PMC article.
-
MITD1 is a brain-specific interferon-inducible factor that inhibits flavivirus replication.Proc Natl Acad Sci U S A. 2025 Mar 25;122(12):e2502064122. doi: 10.1073/pnas.2502064122. Epub 2025 Mar 20. Proc Natl Acad Sci U S A. 2025. PMID: 40112111 Free PMC article.
-
Functional features of a novel interferon-stimulated gene SHFL: a comprehensive review.Front Microbiol. 2023 Dec 11;14:1323231. doi: 10.3389/fmicb.2023.1323231. eCollection 2023. Front Microbiol. 2023. PMID: 38149274 Free PMC article. Review.
-
Serological cross-reactivity among common flaviviruses.Front Cell Infect Microbiol. 2022 Sep 15;12:975398. doi: 10.3389/fcimb.2022.975398. eCollection 2022. Front Cell Infect Microbiol. 2022. PMID: 36189346 Free PMC article. Review.
Cited by
-
The Interactions between Cells and Viruses.Int J Mol Sci. 2024 Jun 23;25(13):6886. doi: 10.3390/ijms25136886. Int J Mol Sci. 2024. PMID: 38999995 Free PMC article.
-
Prolactin Regulates Seasonal Changes in Renal Rheostasis in Djungarian Hamsters.Endocrinology. 2025 Jul 8;166(9):bqaf117. doi: 10.1210/endocr/bqaf117. Endocrinology. 2025. PMID: 40635655 Free PMC article.
-
Genome-wide association analysis of the resistance to infectious hematopoietic necrosis virus in two rainbow trout aquaculture lines confirms oligogenic architecture with several moderate effect quantitative trait loci.Front Genet. 2024 May 24;15:1394656. doi: 10.3389/fgene.2024.1394656. eCollection 2024. Front Genet. 2024. PMID: 38854430 Free PMC article.
-
Omics Integration Uncovers Mechanisms Associated with HIV Viral Load and Potential Therapeutic Insights.medRxiv [Preprint]. 2025 Jul 30:2025.07.29.25332397. doi: 10.1101/2025.07.29.25332397. medRxiv. 2025. PMID: 40766151 Free PMC article. Preprint.
-
Mcl-1 Protein and Viral Infections: A Narrative Review.Int J Mol Sci. 2024 Jan 17;25(2):1138. doi: 10.3390/ijms25021138. Int J Mol Sci. 2024. PMID: 38256213 Free PMC article. Review.
References
-
- HGNC Symbol Report for SHFL. [(accessed on 15 July 2020)]. Available online: https://www.genenames.org/data/gene-symbol-report/#!/hgnc_id/HGNC:2160.
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical

