Genetic and physiological insights into the diazotrophic activity of a non-cyanobacterial marine diazotroph
- PMID: 36302093
- PMCID: PMC10099842
- DOI: 10.1111/1462-2920.16261
Genetic and physiological insights into the diazotrophic activity of a non-cyanobacterial marine diazotroph
Abstract
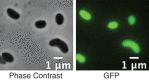
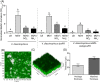
Nitrogen (N2 ) fixation, or diazotrophy, supports a large part of primary production in oceans. Culture-independent approaches highlighted the presence in abundance of marine non-cyanobacterial diazotrophs (NCD), but their ecophysiology remains elusive, mostly because of the low number of isolated NCD and because of the lack of available genetic tools for these isolates. Here, a dual genetic and functional approach allowed unveiling the ecophysiology of a marine NCD affiliated to the species Vibrio diazotrophicus. Physiological characterization of the first marine NCD mutant obtained so far was performed using a soft-gellan assay, demonstrating that a ΔnifH mutant is not able to grow in nitrogen-free media. Furthermore, we demonstrated that V. diazotrophicus produces a thick biofilm under diazotrophic conditions, suggesting biofilm production as an adaptive response of this NCD to cope with the inhibition of nitrogen fixation by molecular oxygen. Finally, the genomic signature of V. diazotrophicus is essentially absent from metagenomic data of Tara Ocean expeditions, despite having been isolated from various marine environments. We think that the genetically tractable V. diazotrophicus strain used in this study may serve as an ideal model to study the ecophysiology of these overlooked procaryotic group.
© 2022 The Authors. Environmental Microbiology published by Applied Microbiology International and John Wiley & Sons Ltd.
Conflict of interest statement
The authors declare that they have no conflict of interest regarding this work.
Figures



Similar articles
-
Non-cyanobacterial diazotrophs: global diversity, distribution, ecophysiology, and activity in marine waters.FEMS Microbiol Rev. 2023 Nov 1;47(6):fuac046. doi: 10.1093/femsre/fuac046. FEMS Microbiol Rev. 2023. PMID: 36416813 Free PMC article. Review.
-
Heterotrophic bacterial diazotrophs are more abundant than their cyanobacterial counterparts in metagenomes covering most of the sunlit ocean.ISME J. 2022 Apr;16(4):927-936. doi: 10.1038/s41396-021-01135-1. Epub 2021 Oct 25. ISME J. 2022. PMID: 34697433 Free PMC article.
-
High metabolic versatility and phenotypic heterogeneity in a marine non-cyanobacterial diazotroph.Curr Biol. 2025 Jun 9;35(11):2659-2671.e3. doi: 10.1016/j.cub.2025.04.071. Epub 2025 May 22. Curr Biol. 2025. PMID: 40409254
-
Global distribution patterns of marine nitrogen-fixers by imaging and molecular methods.Nat Commun. 2021 Jul 6;12(1):4160. doi: 10.1038/s41467-021-24299-y. Nat Commun. 2021. PMID: 34230473 Free PMC article.
-
Marine Non-Cyanobacterial Diazotrophs: Moving beyond Molecular Detection.Trends Microbiol. 2016 Nov;24(11):916-927. doi: 10.1016/j.tim.2016.07.002. Epub 2016 Jul 29. Trends Microbiol. 2016. PMID: 27476748 Review.
Cited by
-
Development and utilization of new O2-independent bioreporters.Microbiol Spectr. 2024 Apr 2;12(4):e0409123. doi: 10.1128/spectrum.04091-23. Epub 2024 Mar 5. Microbiol Spectr. 2024. PMID: 38441526 Free PMC article.
References
-
- Bombar, D. , Paerl, R.W. & Riemann, L. (2016) Marine non‐cyanobacterial diazotrophs: moving beyond molecular detection. Trends in Microbiology, 24, 916–927. - PubMed
-
- Bonaldi, K. , Gargani, D. , Prin, Y. , Fardoux, J. , Gully, D. , Nouwen, N. et al. (2011) Nodulation of Aeschynomene afraspera and A. indica by photosynthetic Bradyrhizobium sp. strain ORS285: the nod‐dependent versus the nod‐independent symbiotic interaction. Molecular Plant‐Microbe Interactions, 24, 1359–1371. - PubMed
-
- Buchfink, B. , Xie, C. & Huson, D.H. (2015) Fast and sensitive protein alignment using DIAMOND. Nature Methods, 12, 59–60. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

