Ubiquitin proteolysis of a CDK-related kinase regulates titan cell formation and virulence in the fungal pathogen Cryptococcus neoformans
- PMID: 36302775
- PMCID: PMC9613880
- DOI: 10.1038/s41467-022-34151-6
Ubiquitin proteolysis of a CDK-related kinase regulates titan cell formation and virulence in the fungal pathogen Cryptococcus neoformans
Abstract
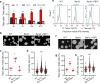
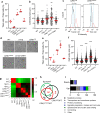
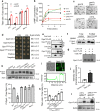
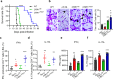
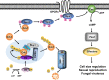
Fungal pathogens often undergo morphological switches, including cell size changes, to adapt to the host environment and cause disease. The pathogenic yeast Cryptococcus neoformans forms so-called 'titan cells' during infection. Titan cells are large, polyploid, display alterations in cell wall and capsule, and are more resistant to phagocytosis and various types of stress. Titan cell formation is regulated by the cAMP/PKA signal pathway, which is stimulated by the protein Gpa1. Here, we show that Gpa1 is activated through phosphorylation by a CDK-related kinase (Crk1), which is targeted for degradation by an E3 ubiquitin ligase (Fbp1). Strains overexpressing CRK1 or an allele lacking a PEST domain exhibit increased production of titan cells similarly to the fbp1∆ mutant. Conversely, CRK1 deletion results in reduced titan cell production, indicating that Crk1 stimulates titan cell formation. Crk1 phosphorylates Gpa1, which then localizes to the plasma membrane and activates the cAMP/PKA signal pathway to induce cell enlargement. Furthermore, titan cell-overproducing strains trigger increased Th1 and Th17 cytokine production in CD4+ T cells and show attenuated virulence in a mouse model of systemic cryptococcosis. Overall, our study provides insights into the regulation of titan cell formation and fungal virulence.
© 2022. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures

Similar articles
-
The Cyclin Cln1 Controls Polyploid Titan Cell Formation following a Stress-Induced G2 Arrest in Cryptococcus.mBio. 2021 Oct 26;12(5):e0250921. doi: 10.1128/mBio.02509-21. Epub 2021 Oct 12. mBio. 2021. PMID: 34634930 Free PMC article.
-
The F-Box protein Fbp1 regulates sexual reproduction and virulence in Cryptococcus neoformans.Eukaryot Cell. 2011 Jun;10(6):791-802. doi: 10.1128/EC.00004-11. Epub 2011 Apr 8. Eukaryot Cell. 2011. PMID: 21478432 Free PMC article.
-
The Transcription Factor Pdr802 Regulates Titan Cell Formation and Pathogenicity of Cryptococcus neoformans.mBio. 2021 Mar 9;12(2):e03457-20. doi: 10.1128/mBio.03457-20. mBio. 2021. PMID: 33688010 Free PMC article.
-
The cAMP/Protein Kinase a Pathway Regulates Virulence and Adaptation to Host Conditions in Cryptococcus neoformans.Front Cell Infect Microbiol. 2019 Jun 18;9:212. doi: 10.3389/fcimb.2019.00212. eCollection 2019. Front Cell Infect Microbiol. 2019. PMID: 31275865 Free PMC article. Review.
-
Cryptococcal Titan Cells: When Yeast Cells Are All Grown up.Curr Top Microbiol Immunol. 2019;422:101-120. doi: 10.1007/82_2018_145. Curr Top Microbiol Immunol. 2019. PMID: 30406867 Review.
Cited by
-
MFPSP: Identification of fungal species-specific phosphorylation site using offspring competition-based genetic algorithm.PLoS Comput Biol. 2024 Nov 18;20(11):e1012607. doi: 10.1371/journal.pcbi.1012607. eCollection 2024 Nov. PLoS Comput Biol. 2024. PMID: 39556608 Free PMC article.
-
The impact of phenotypic heterogeneity on fungal pathogenicity and drug resistance.FEMS Microbiol Rev. 2025 Jan 14;49:fuaf001. doi: 10.1093/femsre/fuaf001. FEMS Microbiol Rev. 2025. PMID: 39809571 Free PMC article. Review.
-
IL-22 and IL-23 regulate the anticryptococcal response during Cryptococcus deuterogattii infection.iScience. 2024 Sep 27;27(10):111054. doi: 10.1016/j.isci.2024.111054. eCollection 2024 Oct 18. iScience. 2024. PMID: 39635124 Free PMC article.
-
Is Cryptococcus neoformans a pleomorphic fungus?Curr Opin Microbiol. 2024 Dec;82:102539. doi: 10.1016/j.mib.2024.102539. Epub 2024 Sep 10. Curr Opin Microbiol. 2024. PMID: 39260180 Free PMC article. Review.
-
Multiple F-Box Proteins Collectively Regulate Cell Development and Pathogenesis in the Human Pathogen Cryptococcus neoformans.J Fungi (Basel). 2022 Nov 29;8(12):1259. doi: 10.3390/jof8121259. J Fungi (Basel). 2022. PMID: 36547592 Free PMC article.
References
-
- Bjorklund M. Cell size homeostasis: metabolic control of growth and cell division. Biochim. Biophys. Acta Mol. Cell Res. 2019;1866:409–417. - PubMed
-
- Lloyd AC. The regulation of cell size. Cell. 2013;154:1194–1205. - PubMed
-
- Odds, F. C. Candida and candidosis: a review and bibliography. (Bailliere Tindall, 1988).
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

