The druggable genome: Twenty years later
- PMID: 36304325
- PMCID: PMC9580872
- DOI: 10.3389/fbinf.2022.958378
The druggable genome: Twenty years later
Abstract
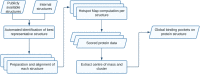
The concept of the druggable genome has been with us for 20 years. During this time, researchers have developed several methods and resources to help assess a target's druggability. In parallel, evidence for target-disease associations has been collated at scale by Open Targets. More recently, the Protein Data Bank in Europe (PDBe) have built a knowledge base matching per-residue annotations with available protein structure. While each resource is useful in isolation, we believe there is enormous potential in bringing all relevant data into a single knowledge graph, from gene-level to protein residue. Automation is vital for the processing and assessment of all available structures. We have developed scalable, automated workflows that provide hotspot-based druggability assessments for all available structures across large numbers of targets. Ultimately, we will run our method at a proteome scale, an ambition made more realistic by the arrival of AlphaFold 2. Bringing together annotations from the residue up to the gene level and building connections within the graph to represent pathways or protein-protein interactions will create complexity that mirrors the biological systems they represent. Such complexity is difficult for the human mind to utilise effectively, particularly at scale. We believe that graph-based AI methods will be able to expertly navigate such a knowledge graph, selecting the targets of the future.
Keywords: artificial intelligence; drug target identification; druggability; druggable genome; knowledge graph; tractability.
Copyright © 2022 Radoux, Vianello, McGreig, Desai and Bradley.
Conflict of interest statement
Authors CR, FV, JM, ND, and AB were employed by the company Exscientia.
Figures
References
LinkOut - more resources
Full Text Sources

