Genetic diversity of Bovine Viral Diarrhea Virus in cattle in France between 2018 and 2020
- PMID: 36304414
- PMCID: PMC9593101
- DOI: 10.3389/fvets.2022.1028866
Genetic diversity of Bovine Viral Diarrhea Virus in cattle in France between 2018 and 2020
Abstract
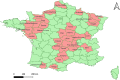
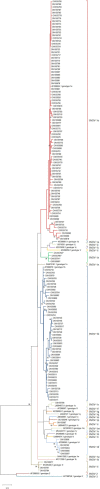
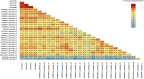
Bovine Viral Diarrhea Virus (BVDV) is one of the main pathogens that affects ruminants worldwide, generating significant economic losses. Like other RNA viruses, BVDV is characterized by a high genetic variability, generating the emergence of new variants, and increasing the risk of new outbreaks. The last report on BVDV genotypes in France was in 2008, since which there have been no new information. The goal of this study is to determine the genetic diversity of BVDV strains currently circulating in France. To this aim, samples of cattle were taken from different departments that are part of the main areas of livestock production during the years 2018 to 2020. Using the partial sequence of the 5'UTR region of the viral genome, we identified and classified 145 samples corresponding to Pestivirus A and one sample corresponding to Pestivirus D. For the Pestivirus A samples, the 1e, 1b, 1d, and 1l genotypes, previously described in France, were identified. Next, the 1r and 1s genotypes, not previously described in the country, were detected. In addition, a new genotype was identified and was tentatively assigned as 1x genotype. These results indicate an increase in the genetic diversity of BVDV in France.
Keywords: Bovine Viral Diarrhea Virus; France; cattle; genetic diversity; genotype.
Copyright © 2022 Rivas, Hasanaj, Deblon, Gisbert and Garigliany.
Conflict of interest statement
Author PG was employed by Ceva Santé Animale. The authors declare that this study received funding from Ceva Santé Animale. The funder was not involved in the laboratory work, analysis and interpretation of the data.
Figures
References
LinkOut - more resources
Full Text Sources

