Identification of energy metabolism-related biomarkers for risk prediction of heart failure patients using random forest algorithm
- PMID: 36304554
- PMCID: PMC9593065
- DOI: 10.3389/fcvm.2022.993142
Identification of energy metabolism-related biomarkers for risk prediction of heart failure patients using random forest algorithm
Abstract
Objective: Energy metabolism plays a crucial role in the improvement of heart dysfunction as well as the development of heart failure (HF). The current study is designed to identify energy metabolism-related diagnostic biomarkers for predicting the risk of HF due to myocardial infarction.
Methods: Transcriptome sequencing data of HF patients and non-heart failure (NF) people (GSE66360 and GSE59867) were obtained from gene expression omnibus (GEO) database. Energy metabolism-related differentially expressed genes (DEGs) were screened between HF and NF samples. The subtyping consistency analysis was performed to enable the samples to be grouped. The immune infiltration level among subtypes was assessed by single sample gene set enrichment analysis (ssGSEA). Random forest algorithm (RF) and support vector machine (SVM) were applied to identify diagnostic biomarkers, and the receiver operating characteristic curves (ROC) was plotted to validate the accuracy. Predictive nomogram was constructed and validated based on the result of the RF. Drug screening and gene-miRNA network were analyzed to predict the energy metabolism-related drugs and potential molecular mechanism.
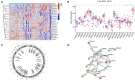
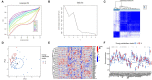
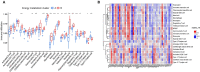
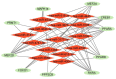
Results: A total of 22 energy metabolism-related DEGs were identified between HF and NF patients. The clustering analysis showed that HF patients could be classified into two subtypes based on the energy metabolism-related genes, and functional analyses demonstrated that the identified DEGs among two clusters were mainly involved in immune response regulating signaling pathway and lipid and atherosclerosis. ssGSEA analysis revealed that there were significant differences in the infiltration levels of immune cells between two subtypes of HF patients. Random-forest and support vector machine algorithm eventually identified ten diagnostic markers (MEF2D, RXRA, PPARA, FOXO1, PPARD, PPP3CB, MAPK14, CREB1, MEF2A, PRMT1) for risk prediction of HF patients, and the proposed nomogram resulted in good predictive performance (GSE66360, AUC = 0.91; GSE59867, AUC = 0.84) and the clinical usefulness in HF patients. More importantly, 10 drugs and 15 miRNA were predicted as drug target and hub miRNA that associated with energy metabolism-related genes, providing further information on clinical HF treatment.
Conclusion: This study identified ten energy metabolism-related diagnostic markers using random forest algorithm, which may help optimize risk stratification and clinical treatment in HF patients.
Keywords: biomarker; energy metabolism; heart failure; nomogram; random forest.
Copyright © 2022 Chen, Jiang, Huang, Chen, Zeng, Wu, Yang, Guo, Li, Wei, Liao, Tse, Sha and Zhuo.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures




References
-
- Zhang F, Zhou G, Guo L, Lu F, Zhou G. Comparison of clinical efficacy of metoprolol combined with irbesartan and hydrochlorothiazide and non-invasive ventilator in the emergency treatment of patients with severe heart failure. Exp Ther Med. (2018) 16:5059–66. 10.3892/etm.2018.6828 - DOI - PMC - PubMed
-
- GBD 2017 Disease and Injury Incidence and Prevalence Collaborators . Global, regional, and national incidence, prevalence, and years lived with disability for 354 diseases and injuries for 195 countries and territories, 1990-2017: a systematic analysis for the Global Burden of Disease Study 2017. Lancet. (2018) 392:1789–858. 10.1016/S0140-6736(18)32279-7 - DOI - PMC - PubMed
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

