RNA binding proteins in MLL-rearranged leukemia
- PMID: 36307883
- PMCID: PMC9615162
- DOI: 10.1186/s40164-022-00343-5
RNA binding proteins in MLL-rearranged leukemia
Abstract
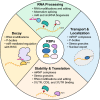
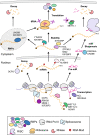
RNA binding proteins (RBPs) have recently emerged as important post-transcriptional gene expression regulators in both normal development and disease. RBPs influence the fate of mRNAs through multiple mechanisms of action such as RNA modifications, alternative splicing, and miR-mediated regulation. This complex and, often, combinatorial regulation by RBPs critically impacts the expression of oncogenic transcripts and, thus, the activation of pathways that drive oncogenesis. Here, we focus on the major features of RBPs, their mechanisms of action, and discuss the current progress in investigating the function of important RBPs in MLL-rearranged leukemia.
Keywords: Leukemia; MLL; RNA binding protein.
© 2022. The Author(s).
Conflict of interest statement
D.S.R is a consultant for AbbVie Inc., providing educational lectures related to clinical work in hematologic malignancies.
Figures


References
-
- Krivtsov AV, Armstrong SA. MLL translocations, histone modifications and leukaemia stem-cell development. Nat Rev Cancer. 2007;7(11):823–833. - PubMed
-
- Moorman AV, Ensor HM, Richards SM, Chilton L, Schwab C, Kinsey SE, et al. Prognostic effect of chromosomal abnormalities in childhood B-cell precursor acute lymphoblastic leukaemia: results from the UK Medical Research Council ALL97/99 randomised trial. Lancet Oncol. 2010;11(5):429–438. - PubMed
-
- Piciocchi A, Messina M, Elia L, Vitale A, Soddu S, Testi AM, et al. Prognostic impact of KMT2A-AFF1-positivity in 926 BCR-ABL1-negative B-lineage acute lymphoblastic leukemia patients treated in GIMEMA clinical trials since 1996. Am J Hematol. 2021;96(9):E334–E338. - PubMed
Publication types
Grants and funding
LinkOut - more resources
Full Text Sources

