Biphasic cell cycle defect causes impaired neurogenesis in down syndrome
- PMID: 36313423
- PMCID: PMC9596798
- DOI: 10.3389/fgene.2022.1007519
Biphasic cell cycle defect causes impaired neurogenesis in down syndrome
Abstract
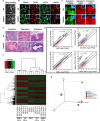
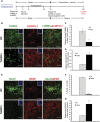
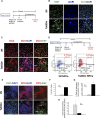
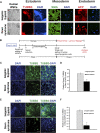
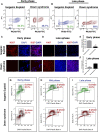
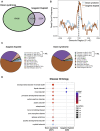
Impaired neurogenesis in Down syndrome (DS) is characterized by reduced neurons, increased glial cells, and delayed cortical lamination. However, the underlying cause for impaired neurogenesis in DS is not clear. Using both human and mouse iPSCs, we demonstrate that DS impaired neurogenesis is due to biphasic cell cycle dysregulation during the generation of neural progenitors from iPSCs named the "neurogenic stage" of neurogenesis. Upon neural induction, DS cells showed reduced proliferation during the early phase followed by increased proliferation in the late phase of the neurogenic stage compared to control cells. While reduced proliferation in the early phase causes reduced neural progenitor pool, increased proliferation in the late phase leads to delayed post mitotic neuron generation in DS. RNAseq analysis of late-phase DS progenitor cells revealed upregulation of S phase-promoting regulators, Notch, Wnt, Interferon pathways, and REST, and downregulation of several genes of the BAF chromatin remodeling complex. NFIB and POU3F4, neurogenic genes activated by the interaction of PAX6 and the BAF complex, were downregulated in DS cells. ChIPseq analysis of late-phase neural progenitors revealed aberrant PAX6 binding with reduced promoter occupancy in DS cells. Together, these data indicate that impaired neurogenesis in DS is due to biphasic cell cycle dysregulation during the neurogenic stage of neurogenesis.
Keywords: Pax6 (RRID: AB_1566562; RRID: AB_2565003; RRID: AB_291612); cell cycle; disease modeling; down syndrome; human iPSCs; mouse iPSCs; neurogenesis; trisomy 21.
Copyright © 2022 Sharma, Nehra, Do, Ghosh, Deshpande and Singhal.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures

Similar articles
-
In Vitro Modeling of Down Syndrome Neurogenesis Using Human-Induced Pluripotent Stem Cells.J Vis Exp. 2025 Mar 7;(217). doi: 10.3791/67382. J Vis Exp. 2025. PMID: 40127053
-
Altered patterning of trisomy 21 interneuron progenitors.Stem Cell Reports. 2022 Jun 14;17(6):1366-1379. doi: 10.1016/j.stemcr.2022.05.001. Epub 2022 May 26. Stem Cell Reports. 2022. PMID: 35623352 Free PMC article.
-
The BAF complex interacts with Pax6 in adult neural progenitors to establish a neurogenic cross-regulatory transcriptional network.Cell Stem Cell. 2013 Oct 3;13(4):403-18. doi: 10.1016/j.stem.2013.07.002. Epub 2013 Aug 8. Cell Stem Cell. 2013. PMID: 23933087 Free PMC article.
-
Neurogenesis impairment: An early developmental defect in Down syndrome.Free Radic Biol Med. 2018 Jan;114:15-32. doi: 10.1016/j.freeradbiomed.2017.07.026. Epub 2017 Jul 27. Free Radic Biol Med. 2018. PMID: 28756311 Review.
-
[Molecular Mechanism Underlying Abnormal Differentiation of Neural Progenitor Cells in the Developing Down Syndrome Brain].Yakugaku Zasshi. 2017;137(7):795-800. doi: 10.1248/yakushi.16-00236-1. Yakugaku Zasshi. 2017. PMID: 28674289 Review. Japanese.
Cited by
-
Transcriptional consequences of trisomy 21 on neural induction.Front Cell Neurosci. 2024 Jan 30;18:1341141. doi: 10.3389/fncel.2024.1341141. eCollection 2024. Front Cell Neurosci. 2024. PMID: 38357436 Free PMC article.
-
Analysis of genotype effects and inter-individual variability in iPSC-derived trisomy 21 neural progenitor cells.Hum Mol Genet. 2025 Jan 23;34(1):85-100. doi: 10.1093/hmg/ddae160. Hum Mol Genet. 2025. PMID: 39533854
-
Extracellular Vesicle Treatment Alleviates Neurodevelopmental and Neurodegenerative Pathology in Cortical Spheroid Model of Down Syndrome.Int J Mol Sci. 2023 Feb 9;24(4):3477. doi: 10.3390/ijms24043477. Int J Mol Sci. 2023. PMID: 36834891 Free PMC article.
-
Consequences of trisomy 21 for brain development in Down syndrome.Nat Rev Neurosci. 2024 Nov;25(11):740-755. doi: 10.1038/s41583-024-00866-2. Epub 2024 Oct 8. Nat Rev Neurosci. 2024. PMID: 39379691 Free PMC article. Review.
References
-
- Aziz N. M., Guedj F., Pennings J. L. A., Olmos-Serrano J. L., Siegel A., Haydar T. F., et al. (2018). Lifespan analysis of brain development, gene expression and behavioral phenotypes in the Ts1Cje, Ts65Dn and Dp(16)1/Yey mouse models of Down syndrome. Dis. Model. Mech. 11 (6), dmm031013. 10.1242/dmm.031013 - DOI - PMC - PubMed
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

