Data Mining Framework for Discovering and Clustering Phenotypes of Atypical Diabetes
- PMID: 36314086
- PMCID: PMC10211492
- DOI: 10.1210/clinem/dgac632
Data Mining Framework for Discovering and Clustering Phenotypes of Atypical Diabetes
Abstract
Context: Some individuals present with forms of diabetes that are "atypical" (AD), which do not conform to typical features of either type 1 diabetes (T1D) or type 2 diabetes (T2D). These forms of AD display a range of phenotypic characteristics that likely reflect different endotypes based on unique etiologies or pathogenic processes.
Objective: To develop an analytical approach to identify and cluster phenotypes of AD.
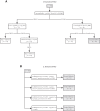
Methods: We developed Discover Atypical Diabetes (DiscoverAD), a data mining framework, to identify and cluster phenotypes of AD. DiscoverAD was trained against characteristics of manually classified patients with AD among 278 adults with diabetes within the Cameron County Hispanic Cohort (CCHC) (Study A). We then tested DiscoverAD in a separate population of 758 multiethnic children with T1D within the Texas Children's Hospital Registry for New-Onset Type 1 Diabetes (TCHRNO-1) (Study B).
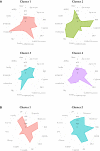
Results: We identified an AD frequency of 11.5% in the CCHC (Study A) and 5.3% in the pediatric TCHRNO-1 (Study B). Cluster analysis identified 4 distinct groups of AD in Study A: cluster 1, positive for the 65 kDa glutamate decarboxylase autoantibody (GAD65Ab), adult-onset, long disease duration, preserved beta-cell function, no insulin treatment; cluster 2, GAD65Ab negative, diagnosed at age ≤21 years; cluster 3, GAD65Ab negative, adult-onset, poor beta-cell function, lacking central obesity; cluster 4, diabetic ketoacidosis (DKA)-prone participants lacking a typical T1D phenotype. Applying DiscoverAD to the pediatric patients with T1D in Study B revealed 2 distinct groups of AD: cluster 1, autoantibody negative, poor beta-cell function, lower body mass index (BMI); cluster 2, autoantibody positive, higher BMI, higher incidence of DKA.
Conclusion: DiscoverAD can be adapted to different datasets to identify and define phenotypes of participants with AD based on available clinical variables.
Keywords: atypical diabetes; bioinformatics; clusters; ketosis-prone diabetes; type 1 diabetes; type 2 diabetes.
© The Author(s) 2022. Published by Oxford University Press on behalf of the Endocrine Society. All rights reserved. For permissions, please e-mail: journals.permissions@oup.com.
Figures

References
-
- Balasubramanyam A. Defining and classifying new subgroups of diabetes. Annu Rev Med. 2021;72:63–74. - PubMed
-
- Balasubramanyam A, Yajnik CS, Tandon N. Non-traditional forms of diabetes worldwide: implications for translational investigation. Transl Endocrinol Metab 2011;2(1):43–67.
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical

