Choosing a cellular model to study SARS-CoV-2
- PMID: 36339347
- PMCID: PMC9634005
- DOI: 10.3389/fcimb.2022.1003608
Choosing a cellular model to study SARS-CoV-2
Abstract
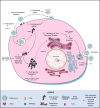
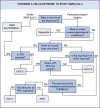
As new pathogens emerge, new challenges must be faced. This is no different in infectious disease research, where identifying the best tools available in laboratories to conduct an investigation can, at least initially, be particularly complicated. However, in the context of an emerging virus, such as SARS-CoV-2, which was recently detected in China and has become a global threat to healthcare systems, developing models of infection and pathogenesis is urgently required. Cell-based approaches are crucial to understanding coronavirus infection biology, growth kinetics, and tropism. Usually, laboratory cell lines are the first line in experimental models to study viral pathogenicity and perform assays aimed at screening antiviral compounds which are efficient at blocking the replication of emerging viruses, saving time and resources, reducing the use of experimental animals. However, determining the ideal cell type can be challenging, especially when several researchers have to adapt their studies to specific requirements. This review strives to guide scientists who are venturing into studying SARS-CoV-2 and help them choose the right cellular models. It revisits basic concepts of virology and presents the currently available in vitro models, their advantages and disadvantages, and the known consequences of each choice.
Keywords: COVID-19; SARS-CoV-2; cell lines; cell model; in vitro approaches; organoids; susceptible cells; viral culture.
Copyright © 2022 Pires De Souza, Le Bideau, Boschi, Wurtz, Colson, Aherfi, Devaux and La Scola.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures


References
-
- Alves E. A., Guimarães A. C. R. (2010). “Cultivo celular,” in Conceitos e métodos para a formação de profissionais em laboratórios de saúde, vol. 2 (Rio de Janeiro: EPSJV; ), 215–252. Available at: http://www.epsjv.fiocruz.br/upload/d/capitulo_5_vol2.pdf.
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Research Materials
Miscellaneous

