Transcription Factor VdCf2 Regulates Growth, Pathogenicity, and the Expression of a Putative Secondary Metabolism Gene Cluster in Verticillium dahliae
- PMID: 36342142
- PMCID: PMC9680623
- DOI: 10.1128/aem.01385-22
Transcription Factor VdCf2 Regulates Growth, Pathogenicity, and the Expression of a Putative Secondary Metabolism Gene Cluster in Verticillium dahliae
Abstract
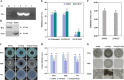
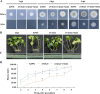
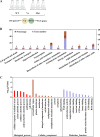
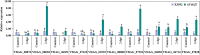
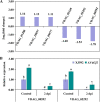
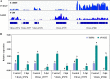
Transcription factors (TFs) bind to the promoters of target genes to regulate gene expression in response to different stimuli. The functions and regulatory mechanisms of transcription factors (TFs) in Verticillium dahliae are, however, still largely unclear. This study showed that a C2H2-type zinc finger TF, VdCf2 (V. dahliae chorion transcription factor 2), plays key roles in V. dahliae growth, melanin production, and virulence. Transcriptome sequencing analysis showed that VdCf2 was involved in the regulation of expression of genes encoding secreted proteins, pathogen-host interaction (PHI) homologs, TFs, and G protein-coupled receptors (GPCRs). Furthermore, VdCf2 positively regulated the expression of VdPevD1 (VDAG_02735), a previously reported virulence factor. VdCf2 thus regulates the expression of several pathogenicity-related genes that also contribute to virulence in V. dahliae. VdCf2 also inhibited the transcription of the Vd276-280 gene cluster and interacted with two members encoding proteins (VDAG_07276 and VDAG_07278) in the gene cluster. IMPORTANCE Verticillium dahliae is an important soilborne phytopathogen which can ruinously attack numerous host plants and cause significant economic losses. Transcription factors (TFs) were reported to be involved in various biological processes, such as hyphal growth and virulence of pathogenic fungi. However, the functions and regulatory mechanisms of TFs in V. dahliae remain largely unclear. In this study, we identified a new transcription factor, VdCf2 (V. dahliae chorion transcription factor 2), based on previous transcriptome data, which participates in growth, melanin production, and virulence of V. dahliae. We provide evidence that VdCf2 regulates the expression of the pathogenicity-related gene VdPevD1 (VDAG_02735) and Vd276-280 gene cluster. VdCf2 also interacts with VDAG_07276 and VDAG_07278 in this gene cluster based on a yeast two-hybrid and bimolecular fluorescence complementation assay. These results revealed the regulatory mechanisms of a pivotal pathogenicity-related transcription factor, VdCf2 in V. dahliae.
Keywords: VdCf2; Verticillium dahliae; gene cluster; pathogenicity; transcription factor; transcriptome.
Conflict of interest statement
The authors declare no conflict of interest.
Figures



References
-
- Agrios GN. 1997. Plant pathology, 4th ed. Academic Press, San Diego, CA.
Publication types
MeSH terms
Substances
Supplementary concepts
LinkOut - more resources
Full Text Sources
Miscellaneous

