A microbial transporter of the dietary antioxidant ergothioneine
- PMID: 36347253
- PMCID: PMC9691600
- DOI: 10.1016/j.cell.2022.10.008
A microbial transporter of the dietary antioxidant ergothioneine
Abstract
Low-molecular-weight (LMW) thiols are small-molecule antioxidants required for the maintenance of intracellular redox homeostasis. However, many host-associated microbes, including the gastric pathogen Helicobacter pylori, unexpectedly lack LMW-thiol biosynthetic pathways. Using reactivity-guided metabolomics, we identified the unusual LMW thiol ergothioneine (EGT) in H. pylori. Dietary EGT accumulates to millimolar levels in human tissues and has been broadly implicated in mitigating disease risk. Although certain microorganisms synthesize EGT, we discovered that H. pylori acquires this LMW thiol from the host environment using a highly selective ATP-binding cassette transporter-EgtUV. EgtUV confers a competitive colonization advantage in vivo and is widely conserved in gastrointestinal microbes. Furthermore, we found that human fecal bacteria metabolize EGT, which may contribute to production of the disease-associated metabolite trimethylamine N-oxide. Collectively, our findings illustrate a previously unappreciated mechanism of microbial redox regulation in the gut and suggest that inter-kingdom competition for dietary EGT may broadly impact human health.
Keywords: ABC transporter; Helicobacter pylori; ergothioneine; host-microbe; low-molecular-weight thiol; metabolism; microbiome; oxidative stress; redox regulation.
Copyright © 2022 Elsevier Inc. All rights reserved.
Conflict of interest statement
Declaration of interests A.S. is a paid consultant of LIV Process.
Figures

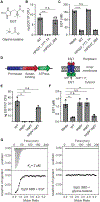





Comment in
-
Bioactive metabolites: The double-edged sword in your food.Cell. 2022 Nov 23;185(24):4469-4471. doi: 10.1016/j.cell.2022.10.022. Cell. 2022. PMID: 36423578
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

