Visualization of G-Quadruplexes, i-Motifs and Their Associates
- PMID: 36348720
- PMCID: PMC9611856
- DOI: 10.32607/actanaturae.11705
Visualization of G-Quadruplexes, i-Motifs and Their Associates
Abstract
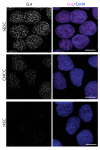
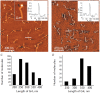
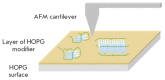
The non-canonical structures formed by G- or C-rich DNA regions, such as quadruplexes and i-motifs, as well as their associates, have recently been attracting increasing attention both because of the arguments in favor of their existence in vivo and their potential application in nanobiotechnology. When studying the structure and properties of non-canonical forms of DNA, as well as when controlling the artificially created architectures based on them, visualization plays an important role. This review analyzes the methods used to visualize quadruplexes, i-motifs, and their associates with high spatial resolution: fluorescence microscopy, transmission electron microscopy (TEM), and atomic force microscopy (AFM). The key approaches to preparing specimens for the visualization of this type of structures are presented. Examples of visualization of non-canonical DNA structures having various morphologies, such as G-wires, G-loops, as well as individual quadruplexes, i-motifs and their associates, are considered. The potential for using AFM for visualizing non-canonical DNA structures is demonstrated.
Keywords: (immuno)fluorescence microscopy; G-quadruplexes; atomic force microscopy; i-motifs; transmission electron microscopy.
Copyright ® 2022 National Research University Higher School of Economics.
Figures







Similar articles
-
The diverse structural landscape of quadruplexes.FEBS Lett. 2019 Aug;593(16):2083-2102. doi: 10.1002/1873-3468.13547. Epub 2019 Jul 30. FEBS Lett. 2019. PMID: 31325371 Review.
-
Direct visualization of G-quadruplexes in DNA using atomic force microscopy.Nucleic Acids Res. 2009 Oct;37(18):6269-75. doi: 10.1093/nar/gkp679. Epub 2009 Aug 20. Nucleic Acids Res. 2009. PMID: 19696072 Free PMC article.
-
RNA G-quadruplexes: G-quadruplexes with "U" turns.Curr Pharm Des. 2012;18(14):2102-11. doi: 10.2174/138161212799958468. Curr Pharm Des. 2012. PMID: 22376111 Review.
-
ElTetrado: a tool for identification and classification of tetrads and quadruplexes.BMC Bioinformatics. 2020 Jan 31;21(1):40. doi: 10.1186/s12859-020-3385-1. BMC Bioinformatics. 2020. PMID: 32005130 Free PMC article.
-
Effect of DNA modifications on the transition between canonical and non-canonical DNA structures in CpG islands during senescence.RSC Adv. 2021 Nov 18;11(59):37205-37217. doi: 10.1039/d1ra07201c. eCollection 2021 Nov 17. RSC Adv. 2021. PMID: 35496393 Free PMC article.
Cited by
-
Artificial Intelligence-Enhanced Analysis of Genomic DNA Visualized with Nanoparticle-Tagged Peptides under Electron Microscopy.Small. 2025 Mar;21(12):e2405065. doi: 10.1002/smll.202405065. Epub 2024 Oct 9. Small. 2025. PMID: 39380435 Free PMC article.
References
LinkOut - more resources
Full Text Sources
Miscellaneous
