Determination of binding affinity of tunicamycin with SARS-CoV-2 proteins: Proteinase, protease, nsp2, nsp9, ORF3a, ORF7a, ORF8, ORF9b, envelope and RBD of spike glycoprotein
- PMID: 36349218
- PMCID: PMC9633632
- DOI: 10.1016/j.vacun.2022.10.006
Determination of binding affinity of tunicamycin with SARS-CoV-2 proteins: Proteinase, protease, nsp2, nsp9, ORF3a, ORF7a, ORF8, ORF9b, envelope and RBD of spike glycoprotein
Abstract
Introduction: Despite the availability of several COVID-19 vaccines, the incidence of infections remains a serious issue. Tunicamycin (TM), an antibiotic, inhibited tumor growth, reduced coronavirus envelope glycoprotein subunit 2 synthesis, and decreased N-linked glycosylation of coronavirus glycoproteins.
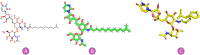
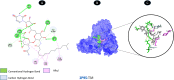
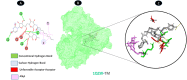
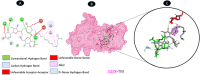
Objectives: Our study aimed to determine how tunicamycin interacts with certain coronavirus proteins (proteinase, protease, nsp9, ORF7a, ORF3a, ORF9b, ORF8, envelope protein, nsp2, and RBD of spike glycoprotein). Methods: Several types of chemo and bioinformatics tools were used to achieve the aim of the study. As a result, virion's effectiveness may be impaired.
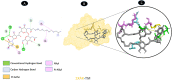
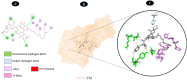
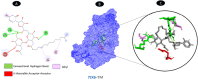
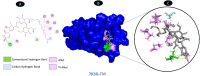
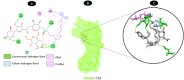
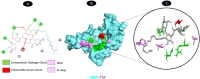
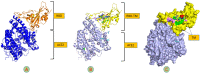
Results: TM can bind to viral proteins with various degrees of affinity. The proteinase had the highest binding affinity with TM. Proteins (ORF9b, ORF8, nsp9, and RBD) were affected by unfavorable donor or acceptor bonds that impact the degree of docking. ORF7a had the weakest affinities.
Conclusions: This antibiotic is likely to effect on SARS-CoV-2 in clinical studies.
Introducción: A pesar de la disponibilidad de varias vacunas contra la COVID-19, la incidencia de infecciones sigue siendo un problema grave. La tunicamicina (TM), un antibiótico, inhibió el crecimiento tumoral, redujo la síntesis de la subunidad 2 de la glicoproteína de la envoltura del coronavirus y disminuyó la glicosilación ligada a N de las glicoproteínas del coronavirus.
Objetivos: Nuestro estudio tuvo como objetivo determinar cómo interactúa la tunicamicina con ciertas proteínas del coronavirus (proteinasa, proteasa, nsp9, ORF7a, ORF3a, ORF9b, ORF8, proteína de la envoltura, nsp2 y RBD de glicoproteína de punta).
Métodos: Se utilizaron varios tipos de herramientas de quimioterapia y bioinformática para lograr el objetivo del estudio. Como resultado, la eficacia del virión puede verse afectada.
Resultados: La TM puede unirse a proteínas virales con diversos grados de afinidad. La proteinasa tenía la mayor afinidad de unión con TM. Las proteínas (ORF9b, ORF8, nsp9 y RBD) se vieron afectadas por enlaces donantes o aceptores desfavorables que afectan el grado de acoplamiento. ORF7a tenía las afinidades más débiles.
Conclusiones: Es probable que este antibiótico tenga efecto sobre el SARS-CoV-2 en estudios clínicos.
Keywords: COVID-19; Docking; Envelope; Spike; Tunicamycin.
© 2022 Elsevier España, S.L.U. All rights reserved.
Conflict of interest statement
There is no conflict of interest in this work.
Figures


Similar articles
-
SARS-CoV-2 Proteins: Are They Useful as Targets for COVID-19 Drugs and Vaccines?Curr Mol Med. 2022;22(1):50-66. doi: 10.2174/1566524021666210223143243. Curr Mol Med. 2022. PMID: 33622224 Review.
-
The role of SARS-CoV-2 accessory proteins in immune evasion.Biomed Pharmacother. 2022 Dec;156:113889. doi: 10.1016/j.biopha.2022.113889. Epub 2022 Oct 17. Biomed Pharmacother. 2022. PMID: 36265309 Free PMC article. Review.
-
Contribution of SARS-CoV-2 Accessory Proteins to Viral Pathogenicity in K18 Human ACE2 Transgenic Mice.J Virol. 2021 Aug 10;95(17):e0040221. doi: 10.1128/JVI.00402-21. Epub 2021 Aug 10. J Virol. 2021. PMID: 34133899 Free PMC article.
-
In silico structural inhibition of ACE-2 binding site of SARS-CoV-2 and SARS-CoV-2 omicron spike protein by lectin antiviral dyad system to treat COVID-19.Drug Dev Ind Pharm. 2022 Oct;48(10):539-551. doi: 10.1080/03639045.2022.2137196. Epub 2022 Oct 27. Drug Dev Ind Pharm. 2022. PMID: 36250723
-
Homology between SARS CoV-2 and human proteins.Sci Rep. 2021 Aug 25;11(1):17199. doi: 10.1038/s41598-021-96233-7. Sci Rep. 2021. PMID: 34433832 Free PMC article.
Cited by
-
Biotransformation of medicarpin from homopterocarpin by Aspergillus niger and its biological characterization.Sci Rep. 2025 Jul 1;15(1):21371. doi: 10.1038/s41598-025-06729-9. Sci Rep. 2025. PMID: 40596118 Free PMC article.
References
-
- Woo P.C., Lau S.K., Laml C.S. Discovery of seven novel mammalian and avian coronaviruses in deltacoronavirus supports bat coronaviruses as the gene source of alphacoronavirus and Betacoronavirus and avian coronaviruses as the gene source of gammacoronavirus and deltacoronavirus. J Virol. 2012;86(7):3995–4008. doi: 10.1128/jvi.06540-11. - DOI - PMC - PubMed
-
- World Health Organization Tracking SARS-CoV-2 variants. 2021. https://www.who.int/en/activities/tracking-SARS-CoV-2-variants/ Accessed 10 February 2022.
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous
