A molecular signature of lung-resident CD8+ T cells elicited by subunit vaccination
- PMID: 36351985
- PMCID: PMC9645351
- DOI: 10.1038/s41598-022-21620-7
A molecular signature of lung-resident CD8+ T cells elicited by subunit vaccination
Abstract
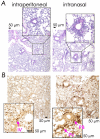
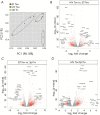
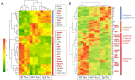
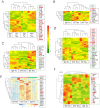
Natural infection as well as vaccination with live or attenuated viruses elicit tissue resident, CD8+ memory T cell (Trm) response. Trm cells so elicited act quickly upon reencounter with the priming agent to protect the host. These Trm cells express a unique molecular signature driven by the master regulators-Runx3 and Hobit. We previously reported that intranasal instillation of a subunit vaccine in a prime boost vaccination regimen installed quick-acting, CD8+ Trm cells in the lungs that protected against lethal vaccinia virus challenge. It remains unexplored whether CD8+ Trm responses so elicited are driven by a similar molecular signature as those elicited by microbes in a real infection or by live, attenuated pathogens in conventional vaccination. We found that distinct molecular signatures distinguished subunit vaccine-elicited lung interstitial CD8+ Trm cells from subunit vaccine-elicited CD8+ effector memory and splenic memory T cells. Nonetheless, the transcriptome signature of subunit vaccine elicited CD8+ Trm resembled those elicited by virus infection or vaccination. Clues to the basis of tissue residence and function of vaccine specific CD8+ Trm cells were found in transcripts that code for chemokines and chemokine receptors, purinergic receptors, and adhesins when compared to CD8+ effector and splenic memory T cells. Our findings inform the utility of protein-based subunit vaccination for installing CD8+ Trm cells in the lungs to protect against respiratory infectious diseases that plague humankind.
© 2022. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures



References
-
- Mackay LK, Rahimpour A, Ma JZ, Collins N, Stock AT, Hafon ML, et al. The developmental pathway for CD103(+)CD8+ tissue-resident memory T cells of skin. Nat. Immunol. 2013;14:1294–1301. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials

