Immune-mediated competition benefits protective microbes over pathogens in a novel host species
- PMID: 36352206
- PMCID: PMC9708653
- DOI: 10.1038/s41437-022-00569-3
Immune-mediated competition benefits protective microbes over pathogens in a novel host species
Abstract
Microbes that protect against infection inhabit hosts across the tree of life. It is unclear whether and how the host immune system may affect the formation of new protective symbioses. We investigated the transcriptomic response of Caenorhabditis elegans following novel interactions with a protective microbe (Enterococcus faecalis) able to defend against infection by pathogenic Staphylococcus aureus. We have previously shown that E. faecalis can directly limit pathogen growth within hosts. In this study, we show that colonisation by protective E. faecalis caused the differential expression of 1,557 genes in pathogen infected hosts, including the upregulation of immune genes such as lysozymes and C-type lectins. The most significantly upregulated host lysozyme gene, lys-7, impacted the competitive abilities of E. faecalis and S. aureus when knocked out. E. faecalis has an increased ability to resist lysozyme activity compared to S. aureus, suggesting that the protective microbe could gain a competitive advantage from this host response. Our finding that protective microbes can benefit from immune-mediated competition after introduction opens up new possibilities for biocontrol design and our understanding of symbiosis evolution. Crosstalk between the host immune response and microbe-mediated protection should favour the continued investment in host immunity and avoid the potentially risky evolution of host dependence.
© 2022. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures


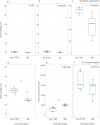
Similar articles
-
Rapid evolution of microbe-mediated protection against pathogens in a worm host.ISME J. 2016 Aug;10(8):1915-24. doi: 10.1038/ismej.2015.259. Epub 2016 Mar 15. ISME J. 2016. PMID: 26978164 Free PMC article.
-
In Vivo Microbial Coevolution Favors Host Protection and Plastic Downregulation of Immunity.Mol Biol Evol. 2021 Apr 13;38(4):1330-1338. doi: 10.1093/molbev/msaa292. Mol Biol Evol. 2021. PMID: 33179739 Free PMC article.
-
Microbe-mediated host defence drives the evolution of reduced pathogen virulence.Nat Commun. 2016 Nov 15;7:13430. doi: 10.1038/ncomms13430. Nat Commun. 2016. PMID: 27845328 Free PMC article.
-
Genetic and molecular analysis of nematode-microbe interactions.Cell Microbiol. 2011 Apr;13(4):497-507. doi: 10.1111/j.1462-5822.2011.01570.x. Epub 2011 Jan 30. Cell Microbiol. 2011. PMID: 21276170 Review.
-
When the microbiome shapes the host: immune evolution implications for infectious disease.Philos Trans R Soc Lond B Biol Sci. 2024 May 6;379(1901):20230061. doi: 10.1098/rstb.2023.0061. Epub 2024 Mar 18. Philos Trans R Soc Lond B Biol Sci. 2024. PMID: 38497259 Free PMC article. Review.
Cited by
-
Characterizing the evolution of defense in a tripartite marine symbiosis using adaptive dynamics.Evol Lett. 2024 Oct 20;9(1):105-114. doi: 10.1093/evlett/qrae052. eCollection 2025 Feb. Evol Lett. 2024. PMID: 39906587 Free PMC article.
-
Distinct members of the Caenorhabditis elegans CeMbio reference microbiota exert cryptic virulence that is masked by host defense.Mol Microbiol. 2024 Sep;122(3):387-402. doi: 10.1111/mmi.15258. Epub 2024 Apr 16. Mol Microbiol. 2024. PMID: 38623070
-
Lysozyme Inhibitors as Tools for Lysozyme Profiling: Identification and Antibacterial Function of Lysozymes in the Hemolymph of the Blue Mussel.Molecules. 2023 Oct 13;28(20):7071. doi: 10.3390/molecules28207071. Molecules. 2023. PMID: 37894549 Free PMC article.
-
The role of animal hosts in shaping gut microbiome variation.Philos Trans R Soc Lond B Biol Sci. 2024 May 6;379(1901):20230071. doi: 10.1098/rstb.2023.0071. Epub 2024 Mar 18. Philos Trans R Soc Lond B Biol Sci. 2024. PMID: 38497257 Free PMC article. Review.
-
Tolerance-conferring defensive symbionts and the evolution of parasite virulence.Evol Lett. 2023 May 5;7(4):262-272. doi: 10.1093/evlett/qrad015. eCollection 2023 Aug. Evol Lett. 2023. PMID: 37475754 Free PMC article.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

