MXene-Based Nucleic Acid Biosensors for Agricultural and Food Systems
- PMID: 36354491
- PMCID: PMC9688781
- DOI: 10.3390/bios12110982
MXene-Based Nucleic Acid Biosensors for Agricultural and Food Systems
Abstract
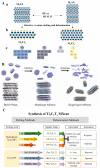
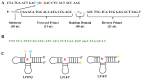
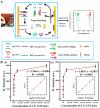
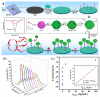
MXene is a two-dimensional (2D) nanomaterial that exhibits several superior properties suitable for fabricating biosensors. Likewise, the nucleic acid (NA) in oligomerization forms possesses highly specific biorecognition ability and other features amenable to biosensing. Hence the combined use of MXene and NA is becoming increasingly common in biosensor design and development. In this review, MXene- and NA-based biosensors are discussed in terms of their sensing mechanisms and fabrication details. MXenes are introduced from their definition and synthesis process to their characterization followed by their use in NA-mediated biosensor fabrication. The emphasis is placed on the detection of various targets relevant to agricultural and food systems, including microbial pathogens, chemical toxicants, heavy metals, organic pollutants, etc. Finally, current challenges and future perspectives are presented with an eye toward the development of advanced biosensors with improved detection performance.
Keywords: MXene; agricultural and food system; biosensors; contaminants; nucleic acid.
Conflict of interest statement
The authors declare no conflict of interest.
Figures






Similar articles
-
MXene-based aptasensor for the detection of aflatoxin in food and agricultural products.Environ Pollut. 2023 Jan 1;316(Pt 1):120695. doi: 10.1016/j.envpol.2022.120695. Epub 2022 Nov 21. Environ Pollut. 2023. PMID: 36423887 Review.
-
Recent Advancements in MXene-Based Biosensors for Health and Environmental Applications-A Review.Biosensors (Basel). 2024 Oct 12;14(10):497. doi: 10.3390/bios14100497. Biosensors (Basel). 2024. PMID: 39451710 Free PMC article. Review.
-
Recent advances on functional nucleic acid-based biosensors for detection of food contaminants.Talanta. 2021 Jan 15;222:121565. doi: 10.1016/j.talanta.2020.121565. Epub 2020 Aug 26. Talanta. 2021. PMID: 33167261 Review.
-
MXene-Based Electrochemical Biosensors: Advancing Detection Strategies for Biosensing (2020-2024).Biosensors (Basel). 2025 Feb 20;15(3):127. doi: 10.3390/bios15030127. Biosensors (Basel). 2025. PMID: 40136924 Free PMC article. Review.
-
Recent Advances in MXene Nanocomposite-Based Biosensors.Biosensors (Basel). 2020 Nov 20;10(11):185. doi: 10.3390/bios10110185. Biosensors (Basel). 2020. PMID: 33233574 Free PMC article. Review.
Cited by
-
Gold nanoparticles-doped MXene heterostructure for ultrasensitive electrochemical detection of fumonisin B1 and ampicillin.Mikrochim Acta. 2024 May 2;191(5):294. doi: 10.1007/s00604-024-06369-2. Mikrochim Acta. 2024. PMID: 38698253
References
-
- Manzanares-Palenzuela C.L., Martín-Fernández B., López M.S.-P., López-Ruiz B. Electrochemical genosensors as innovative tools for detection of genetically modified organisms. TrAC Trends Anal. Chem. 2015;66:19–31. doi: 10.1016/j.trac.2014.10.006. - DOI
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

