Selective PROTAC-mediated degradation of SMARCA2 is efficacious in SMARCA4 mutant cancers
- PMID: 36357397
- PMCID: PMC9649729
- DOI: 10.1038/s41467-022-34562-5
Selective PROTAC-mediated degradation of SMARCA2 is efficacious in SMARCA4 mutant cancers
Abstract
The mammalian SWItch/Sucrose Non-Fermentable (SWI/SNF) helicase SMARCA4 is frequently mutated in cancer and inactivation results in a cellular dependence on its paralog, SMARCA2, thus making SMARCA2 an attractive synthetic lethal target. However, published data indicates that achieving a high degree of selective SMARCA2 inhibition is likely essential to afford an acceptable therapeutic index, and realizing this objective is challenging due to the homology with the SMARCA4 paralog. Herein we report the discovery of a potent and selective SMARCA2 proteolysis-targeting chimera molecule (PROTAC), A947. Selective SMARCA2 degradation is achieved in the absence of selective SMARCA2/4 PROTAC binding and translates to potent in vitro growth inhibition and in vivo efficacy in SMARCA4 mutant models, compared to wild type models. Global ubiquitin mapping and proteome profiling reveal no unexpected off-target degradation related to A947 treatment. Our study thus highlights the ability to transform a non-selective SMARCA2/4-binding ligand into a selective and efficacious in vivo SMARCA2-targeting PROTAC, and thereby provides a potential new therapeutic opportunity for patients whose tumors contain SMARCA4 mutations.
© 2022. The Author(s).
Conflict of interest statement
Authors are employees of Genentech and/or Arvinas (as defined in affiliations) and own shares of Roche or Arvinas stock, respectively. The authors declare no competing interests.
Figures

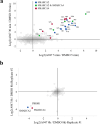



References
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Miscellaneous

